ForliLab
448 posts


ForliLab
@ForliLab
Molecular Modeling & Drug Design. The lab of Stefano Forli at Dept. of Integrative Structural & Computational Biology @scrippsresearch Home of the AutoDock
La Jolla, CA (USA) انضم Eylül 2018
378 يتبع948 المتابعون
ForliLab أُعيد تغريده

Transthyretin’s structural asymmetry & “molecular breathing” drive its misfolding in ATTR—a discovery by Prof. Gabriel Lander (@LanderLab) & Prof. Jeff Kelly, published in @NatureSMB, that could inform new therapies for amyloid diseases. More: ow.ly/xCx850UVbbl

English

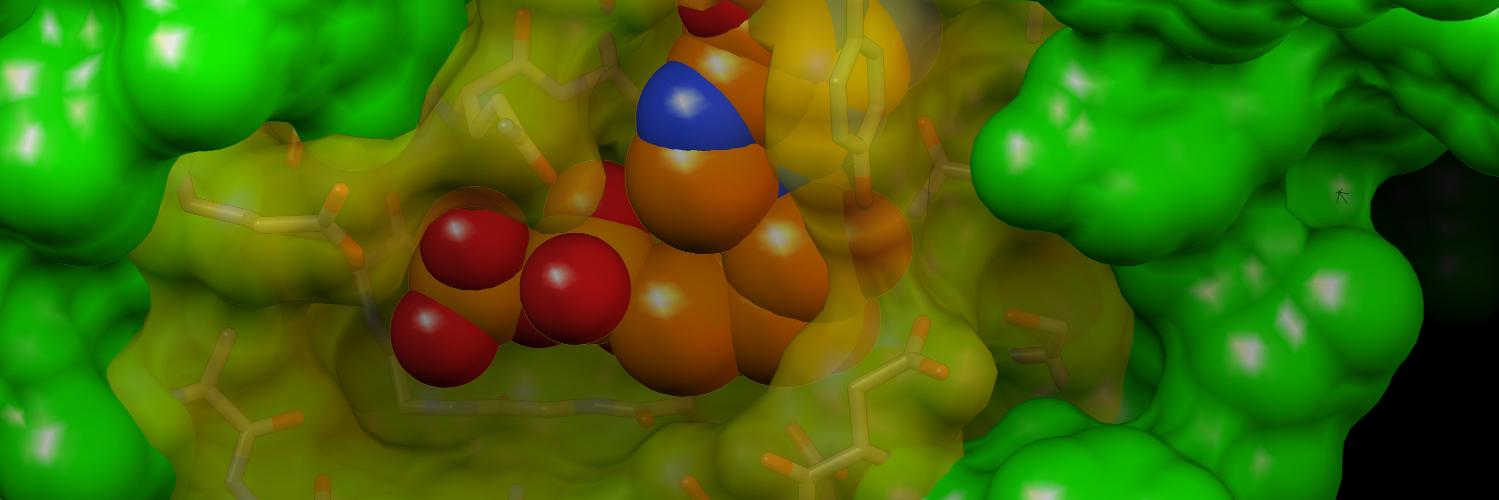
Excited to announce the publication of our new paper on HIV-1 capsid stability! cell.com/biophysj/fullt… #HIV #MolecularDynamics
English
ForliLab أُعيد تغريده

Kudos @ForliLab!!!
JCIM & JCTC Journals@JCIM_JCTC
#HighlightOfTheWeek Cosolvent molecular dynamics are an increasingly popular form of MD simulations where small molecule cosolvents are added to the system, helping to identify cryptic and allosteric pockets in proteins. #compchem pubs.acs.org/doi/10.1021/ac…
Español

Our pre-print with full method and validation details is also available on ChemRxiv: chemrxiv.org/engage/chemrxi… (4/4)
English

CryoXKit uses ligand density from x-ray crystallography and/or cryo-EM to guide dockings with AutoDock, resulting in better pose prediction and improved virtual screening performance across multiple systems. You can find the new tool now on GitHub: github.com/forlilab/CryoX…
(3/4)
English
ForliLab أُعيد تغريده

If you’re interested in fast sampling and flexible and automated creation of cosolvent systems with @openmm_toolkit check this out! Huge thanks to the team @je_eberhardt @diogo_stmart @manuco17 @moonii1020 @Joe_r_loeffler @ForliLab
pubs.acs.org/doi/10.1021/ac… ⚛️🧬
English

Ringtail is available as an open-source package on PyPi, conda-forge, or by downloading and installing source code directly from GitHub (github.com/forlilab/Ringt…). Installation instructions, usage examples, and extensive documentation are available here ringtail.readthedocs.io/en/latest/
English

Buried under a pile of docking data and not sure where to go next? Ringtail 2 will help you organize, filter, and visualize docking data via an easy-to-use command line tool, or integrated in your #python scripts using the brand new API! #autodockgpu #autodockvina

English
ForliLab أُعيد تغريده

AlphaFold2+MD simulations = powerful insights into protein conformational ensembles, but can it capture all experimental structural variations? Great collaborative effort!! @Structure_CP @RRiccabona @OPIGlets @scrippsresearch @DTUtweet #compchem sciencedirect.com/science/articl…
English
ForliLab أُعيد تغريده
Mesoscale Explorer molstar.org/me/ Paper out !! Excellent work with @asrmoin @dsgoodsell and @sehnaldavid
doi.org/10.1002/pro.51…
English

Our second years @AllisonBarkdull, @YutingZhan8328, and @Renhao4 received lab coats this weekend! #computationalscientists #whataretheygonnadowithlabcoats

English

congratulations to Ishan for winning second place for his talk at Scripps' Graduate Research Symposium #IshanFold

English

Do you know Python and have experience with RDKit?
Do you want to work in an exciting and multidisciplinary environment?The AutoDock project is hiring a programmer, apply here! recruiting2.ultipro.com/SCR1003TSRI/Jo…
English



