Dave Rhee retweetet
Dave Rhee
146 posts


Dave Rhee
@DaveYRhee
Dad | Molecular Glues | Proteomics
Boston, MA Beigetreten Nisan 2013
733 Folgt146 Follower
Dave Rhee retweetet

🧩 New paper out in Nature Communications!
"Unveiling the hidden interactome of CRBN molecular glues"
Huge thanks to our incredible team and collaborators who made this possible!
📖 Read the full open-access article here: nature.com/articles/s4146…
English
Dave Rhee retweetet
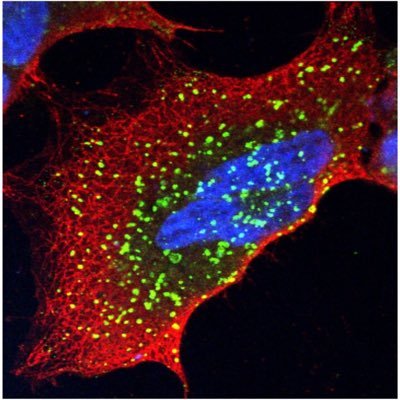
EndoMAP.v1 charts the structural landscape of human early endosome complexes
nature.com/articles/s4158…
English
Dave Rhee retweetet

Check out our latest study @NatureSMB: Establishing a consensus model for #ubiquitin chain assembly by HECT #E3 ligases:#cryoEM structures of #TRIP12 forming K29-linked and K29/K48-branched chains!
❕Publication: nature.com/articles/s4159…
@SamuelMaiwald @UniLeiden
English
Dave Rhee retweetet

#Technology Chemical tools to expand the ligandable proteome: Diversity-oriented synthesis-based photoreactive stereoprobes by @d_ogasa, @brumelillo, @bencravatt at @scrippsresearch @broadinstitute @harvardmed @MassGeneralNews dlvr.it/TGwBK2
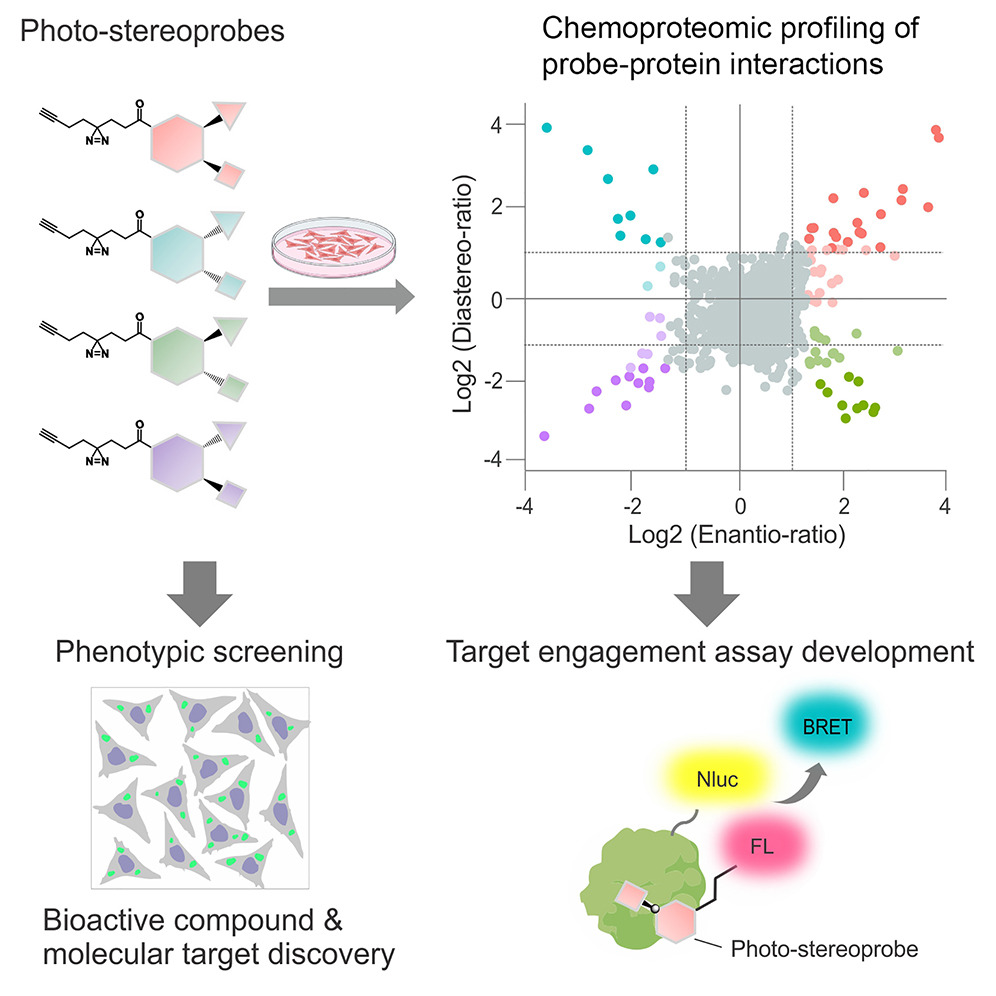
English
Dave Rhee retweetet
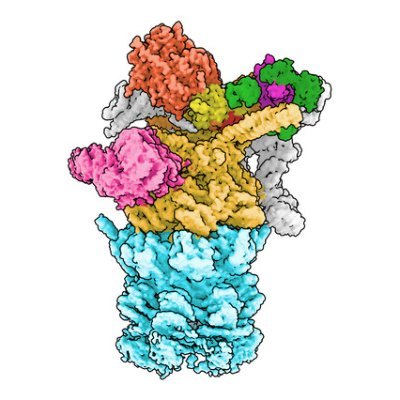
We are very excited to share our recent preprint on the nuclear import mechanism of the proteasome, driven by the multivalent adaptor AKIRIN2!
biorxiv.org/content/10.110… 🧵 (1/7)
English
Dave Rhee retweetet

Very excited to present the structural and mechanistic basis for stress response silencing by the monster E3 ligase SIFI.
biorxiv.org/content/10.110…
English
Dave Rhee retweetet

Thrilled to see our work out in @naturemethods today! We applied prime editing to multiplexed dropout screens and benchmarked the tool’s ability to induce phenotypes when installing premature stop codons in essential genes.
Nature Methods@naturemethods
From @bsadamson and colleagues, a prime editing platform for high-throughput functional interrogation of up to tens of thousands of small genetic variants. nature.com/articles/s4159…
English
Dave Rhee retweetet

Online now! Chemical tools to expand the ligandable proteome: Diversity-oriented synthesis-based photoreactive stereoprobes by @d_ogasa, @brumelillo, @bencravatt, et al at @scrippsresearch @broadinstitute @harvardmed dlvr.it/TGFYHz
English
Dave Rhee retweetet

Structural basis for C-degron selectivity across KLHDCX family E3 ubiquitin ligases, now out @SpringerNature @NatureComms
Awesome collaboration with @sagarchittori .
@StJudeStructBio @MPI_Biochem @StJudeResearch #TPD #PROTAC #SchulmanLab
rdcu.be/d0h2G
English
Dave Rhee retweetet

6 years of work +
Over 2 years in review +
1 extremely talented graduate student @LeonaNease +
Amazing collaborators =
One extremely cool new mechanism that regulates metastasis through modification of the selenocysteine tRNA!
nature.com/articles/s4301…
English
Dave Rhee retweetet

Check out our recent Opinion in @TrendsinPharma on chemical proteomic binding site determination for non-covalent compounds
Trends in Pharmacological Sciences@TrendsinPharma
Chemical proteomic mapping of reversible small molecule binding sites in native systems dlvr.it/TFMRnc
English
Dave Rhee retweetet

Postdoctoral Position in @businolab
Come join us as a #postdoc studying the mechanisms by which the ubiquitin-proteasome system (UPS) controls cancer.
jobrxiv.org/job/dr-luca-bu…
English
Dave Rhee retweetet

Open season on FOXA1! Our paper with the Cravatt group is out today in @MolecularCell. Please check out the updated data, including a new discovery that FOXA1 ligands mirror the effects of a nearby prostate cancer mutation. authors.elsevier.com/a/1jxLO3vVUPRn… (1/2)


English
Dave Rhee retweetet

Principles of paralog-specific degradation engaging the C-degron E3 KLHDC2, now out @SpringerNature @NatureComms
Awesome collaboration with @Richard_E_Lee and Taosheng Chen labs.
@StJudeStructBio @MPI_Biochem @StJudeResearch #TPD #PROTAC #SchulmanLab
rdcu.be/dWKpF
English
Dave Rhee retweetet

Thrilled to share our latest preprint describing discovery of the charged, metabolically activated molecular glue degrader engaging the basic pocket of the structural CRBN homolog, YPEL5 - a subunit of the giant GID/CTLH E3 ligase!!!
biorxiv.org/content/10.110…
English

Excited to join the @HHMINEWS community - made possible by talented lab members, current and former, and amazing colleagues at @HarvardCellBio and beyond!
English
Dave Rhee retweetet

Excited to share our new paper @PLOSGenetics that reveals how Shh signaling promotes myosin II activity to regulate tooth morphogenesis in mice! doi.org/10.1371/journa…
English
Dave Rhee retweetet

Very pleased to share our latest work on substrate identification of deubiquitylases, led by @VRossio, in collaboration with @PauloLabMS @GygiLab, available @CellChemBiol. Link below is active until July 17. authors.elsevier.com/c/1jA778jWWJxL…
English


