Bioecomatics
5.6K posts

Bioecomatics
@bioecomatic
Applied BiomedicoClinical Informatics BioSystemsInternalMedicine CM2EnvM,OrganSys2Epigenetics,NueOx(aVIZaid2NuerOncoSTx) PsySEMECEpi/Bioecological LangofOrgeff
Tokyo-to, Japan Se unió Şubat 2024
1.6K Siguiendo113 Seguidores
Tweet fijado
Bioecomatics retuiteado

Gratitude rewires the brain.
Gratitude rewires the brain.
Gratitude rewires the brain.
Gratitude rewires the brain.
Gratitude rewires the brain.
Gratitude rewires the brain.
Gratitude rewires the brain.
Gratitude rewires the brain.

Curiosity@CuriosityonX
🚨: Every time you express gratitude, your brain physically rewrites itself, making you naturally more positive and resilient, neurology says.
English
Bioecomatics retuiteado
Bioecomatics retuiteado
Bioecomatics retuiteado

Wearable devices and cardiovascular health: revolutionizing remote monitoring and disease prevention. Read this State-of-the-Art review just published in #EHJ
👉 ow.ly/nZX450YAtq0
@RoccoMontone @ehj_ed #prevention

English
Bioecomatics retuiteado
Bioecomatics retuiteado

Bioecomatics retuiteado

AlphaFold Database expands to proteome-scale quaternary structures
1 AFDB is extended beyond monomers by adding 1,754,242 high-confidence predicted protein complexes (primarily homodimers), enabling proteome-scale access to quaternary structure models with standardized formats, metrics, and metadata for search/visualization/bulk use.
2 The study runs a large-scale prediction campaign covering ~31M complexes from 4,777 proteomes: 23,441,822 homodimers derived from UniProt proteomes, plus 7,620,644 heterodimer candidates derived from STRING “physical interaction” links for 16 model organisms and 30 WHO-prioritized global health proteomes.
3 A key methodological contribution is confidence calibration for complex models using interface-focused signals. The authors evaluate multiple metrics and settle on a practical high-confidence rule: ipSAEmin ≥ 0.6, pLDDTavg ≥ 70, and backbone clashes ≤ 10, benchmarked against post-2021 PDB homodimers (positives) and PDB monomers (negatives).
4 Among interface metrics tested (ipTM, ipSAEmin, LISmin, pDockQ2), ipSAEmin provides the clearest separation between true homodimers and monomers and shows a stable F1 plateau up to the chosen cutoff (precision ~0.859, recall ~0.655, F1 ~0.744), motivating its use as the primary AFDB-facing filter.
5 The resulting high-confidence set retains ~7% of predicted homodimers (~1.8M out of ~23M), and AFDB further labels entries by ipSAEmin into “very high-confidence” (≥0.8), “confident” (0.7–<0.8), and “low-confidence” (0.6–<0.7) to help non-expert users interpret interfaces.
6 Compared to experimentally determined multimers in the PDB, the high-confidence predicted complexes increase structural coverage by 1–3 orders of magnitude for most organisms, with especially large gaps being bridged in Metazoa and Viridiplantae (exceptions are heavily studied species like human, E. coli, and yeast).
7 Dataset consistency is assessed at scale: after clustering sequences at 98% identity/95% coverage and aligning complexes within clusters, 95.9% of aligned complexes show complex TM-score > 0.8; additionally, chain A vs chain B within homodimers yields 98.81% with TM-score > 0.8, supporting internal coherence of the predictions.
8 Prediction yield varies across taxonomy: archaea and bacteria show >3× higher high-confidence homodimer rates than eukaryotes, consistent with shorter/more compact prokaryotic proteins and a higher prevalence of homo-oligomeric assemblies, while eukaryotic proteins are often longer, multi-domain, and more disordered and may preferentially form heteromers.
9 For heterodimers, applying the same thresholds produces 56,956 “tentatively high-confidence” models from 7,620,644 STRING-derived candidates. High-confidence likelihood correlates with STRING score, but also with homodimer-like properties (higher inter-chain sequence identity and smaller chain-length differences), motivating further calibration tailored to heteromer biology.
10 Structural clustering of 1,811,201 predicted complexes (high-confidence homodimers + tentative heterodimers) compresses the space ~8-fold into 224,862 clusters; complex topology is highly recurrent (top 1% of non-singleton representatives cover ~25% of entries; top 20% cover ~82%), and ~9% of non-singleton clusters span multiple superkingdoms, suggesting conserved complex building blocks across deep evolution.
11 Case studies highlight “emergent” structures only visible in oligomeric context: a domain-swapped fold in Dictyostelium Q55DI5 becomes high-confidence only as a dimer; a fungal membrane protein (Atg33) gains a more coherent assembly and clearer membrane boundaries as a dimer; multimer prediction can also refine inter-domain architecture even when monomer predictions are already confident, and can partially rescue low-confidence monomers (e.g., an HTH regulator consistent with dimeric function).
12 Engineering/scaling details matter for feasibility at this size: MMseqs2-GPU MSAs (best hit per taxon as an orthology-like filter), AlphaFold-Multimer inference via accelerated ColabFold/OpenFold (TensorRT + cuEquivariance), batching strategies to reduce recompilations, and a post-processing pipeline producing AFDB-compliant mmCIF/BCIF plus computed interface metrics and clash scores.
📜Paper:
biorxiv.org/content/10.648…
#AlphaFold #ProteinComplexes #StructuralBioinformatics #Interactome #ComputationalBiology #Bioinformatics #ProteinStructure #AIforScience #OpenScience

English
Bioecomatics retuiteado

The idea that the universe is fundamentally mathematical is one of the leading frameworks in modern thought. | iai.tv/articles/mathe…
But physicist Martín López Corredoira invites us to reconsider, questioning whether mathematics describes the world as it is, or as we are able to conceive it.

English
Bioecomatics retuiteado

Grand Challenges for Predictive Modeling in Small Molecule Drug Discovery
chemrxiv.org/doi/full/10.26…
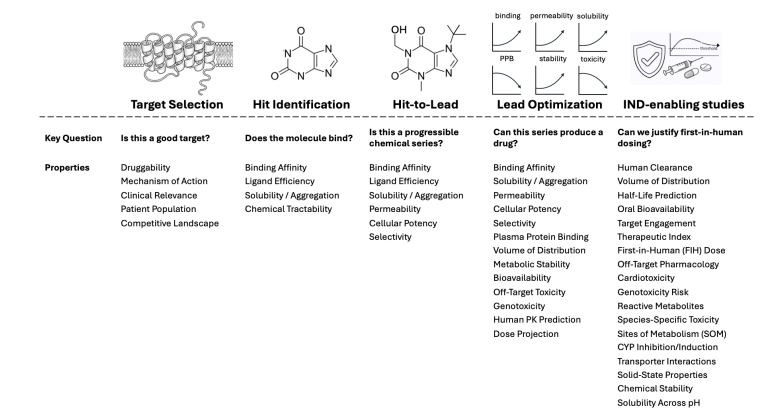
In the study. authors defines a set of Grand Challenges for predictive modeling in small-molecule drug discovery, aiming to clarify where computational methods can most meaningfully advance real drug discovery. Rather than providing a broad methodological review, the paper identifies specific scientific and technical barriers that currently limit progress and proposes measurable benchmarks to evaluate future advances. The goal is to align researchers, industry leaders, and investors around problems that have direct impact on drug discovery outcomes.
The challenges are organized into four core domains: Chemistry, Structure, Energy, and Pharmacology. For each domain, the authors define key problems, describe the underlying physical and biological principles, explain their relevance to drug discovery decision-making, assess the current state of the field, and propose quantitative metrics for measuring progress. By focusing on well-defined problems rather than general technological trends, the work aims to bridge the gap between rapidly advancing computational tools (including AI, machine learning, robotics, high-performance computing, and quantum computing) and the practical needs of medicinal chemistry and pharmacology. The overarching objective is to catalyze measurable, sustained progress in computational drug discovery rather than short-lived advances driven by enthusiasm or benchmarking alone.
#BioAI #AI_drug_discovery
English
Bioecomatics retuiteado

Artificial Biological Intelligence (ABI)
In a post-Darwinian era of being able to write genomes, the implications—both for good and harm—are profound. In conversation with @AdrianWoolfson on his new book On the Future of Species
erictopol.substack.com/p/on-the-futur…

English
Bioecomatics retuiteado

Mathematical Methods in Data Science — Bridging Theory and Applications with Python: amzn.to/4b7ZYQ4
——————
#ML #MachineLearning #DataScientist #DataScience #Mathematics #Algorithms

English
Bioecomatics retuiteado

Mortality from #cardiovascular disease and #diabetes has declined among people with diabetes in most jurisdictions, whereas mortality from #dementia has increased markedly, independent of age thelancet.com/journals/landi…
#T2D #CVD
English
Bioecomatics retuiteado

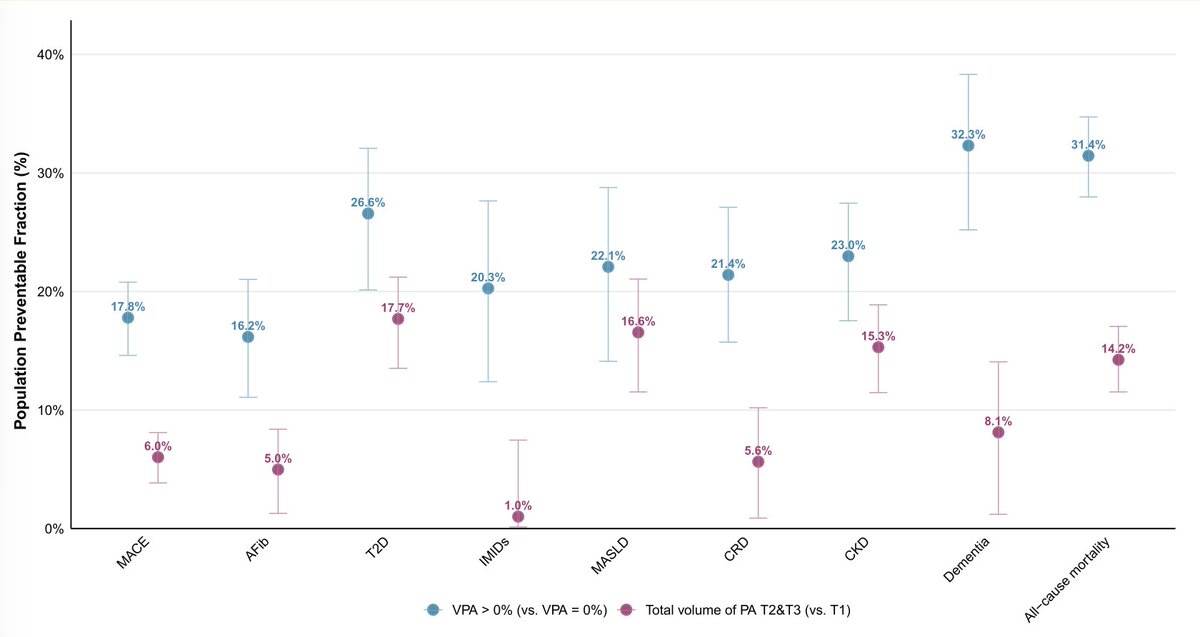
Intensity of exercise vs volume of physical activity made a difference for lower risks of 8 diseases and all-cause mortality among 96,000 @uk_biobank participants, especially noted for immune-mediated (IMID). VPA-vigorous physical activity
academic.oup.com/eurheartj/adva…
English
Bioecomatics retuiteado

Bioecomatics retuiteado
Bioecomatics retuiteado
Bioecomatics retuiteado

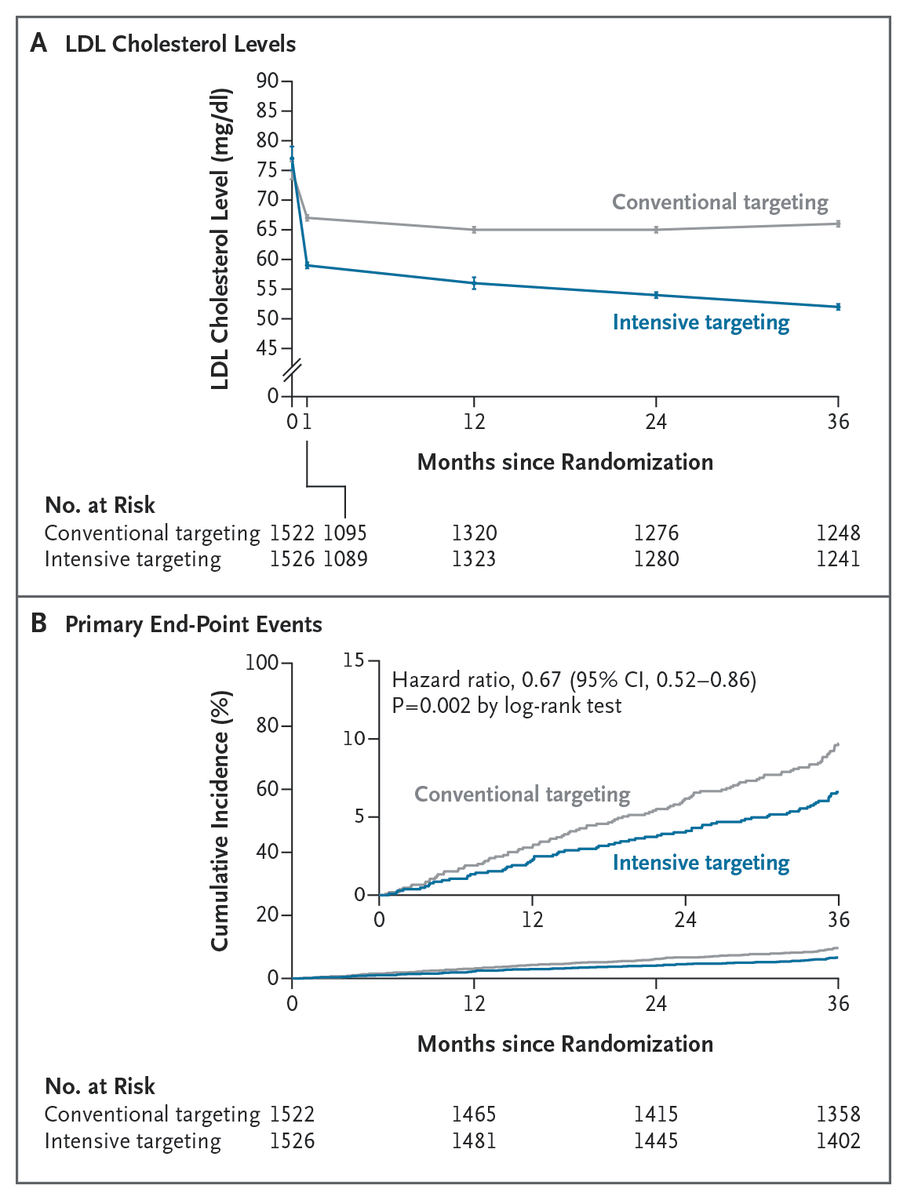
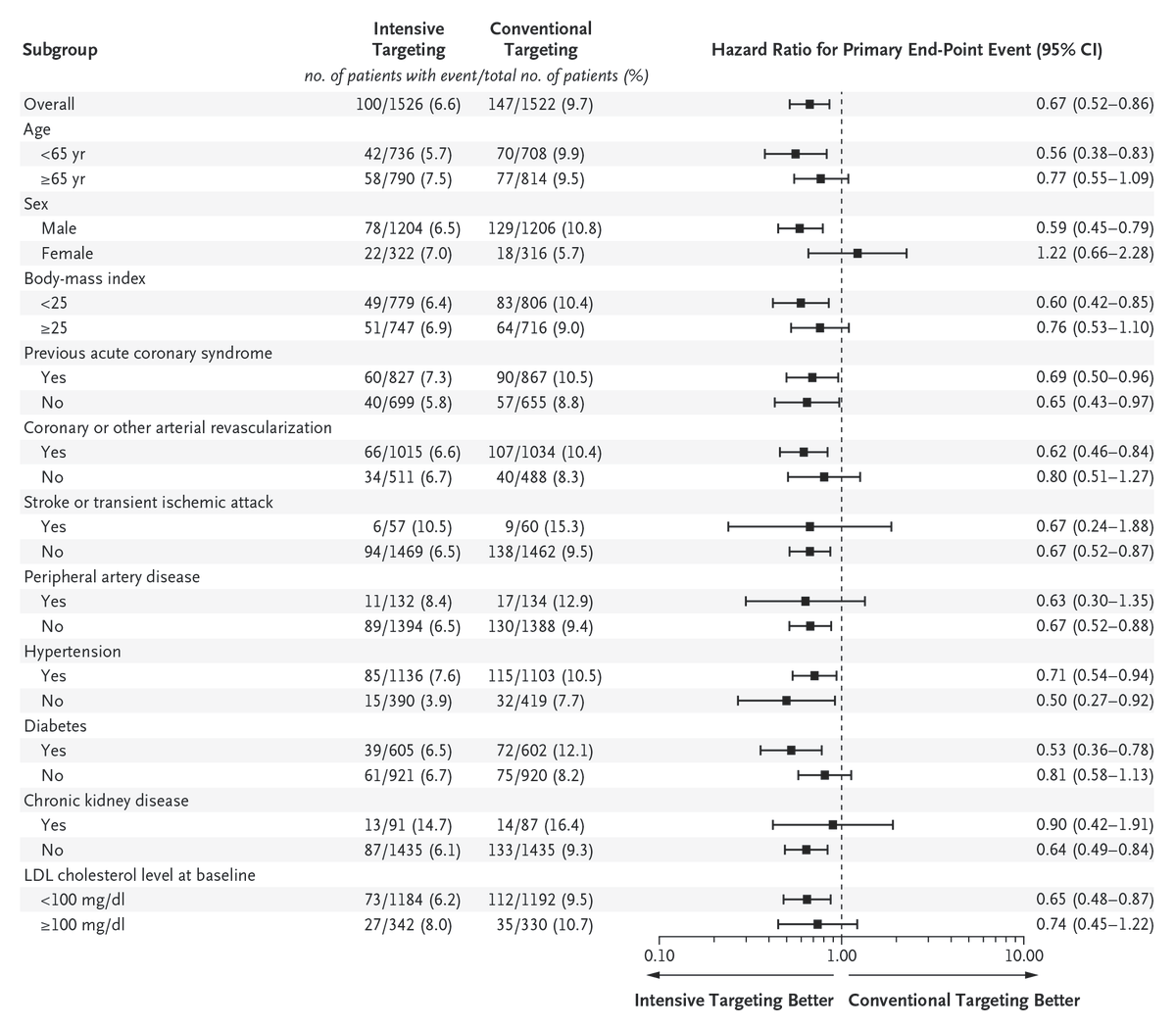
Original Article: Intensive LDL Cholesterol Targeting in Atherosclerotic Cardiovascular Disease (Ez-PAVE trial) nejm.org/doi/full/10.10…
Editorial: Paving the Road toward Targeted Lipid Lowering nejm.org/doi/full/10.10…
#ACC26 | @ACCinTouch
English
Bioecomatics retuiteado

Bioecomatics retuiteado

Presented at #ACC26:
Among patients with atherosclerotic cardiovascular disease, targeting an LDL cholesterol level below 55 mg per deciliter led to a lower 3-year risk of cardiovascular events than targeting a level below 70 mg per deciliter. Full Ez-PAVE trial results: nejm.org/doi/full/10.10…
Editorial: Paving the Road toward Targeted Lipid Lowering nejm.org/doi/full/10.10…
@ACCinTouch

English