Alexander W. Dowling
84 posts


Alexander W. Dowling
@DowlingLab
Associate Professor in @NDCBE at @NotreDame. Process Systems Engineering. Computational Optimization. Data Science. Statistics.





University of Notre Dame ranks as top educational institution and in top 20 on Forbes’ America’s Best Large Employers list news.nd.edu/news/universit…
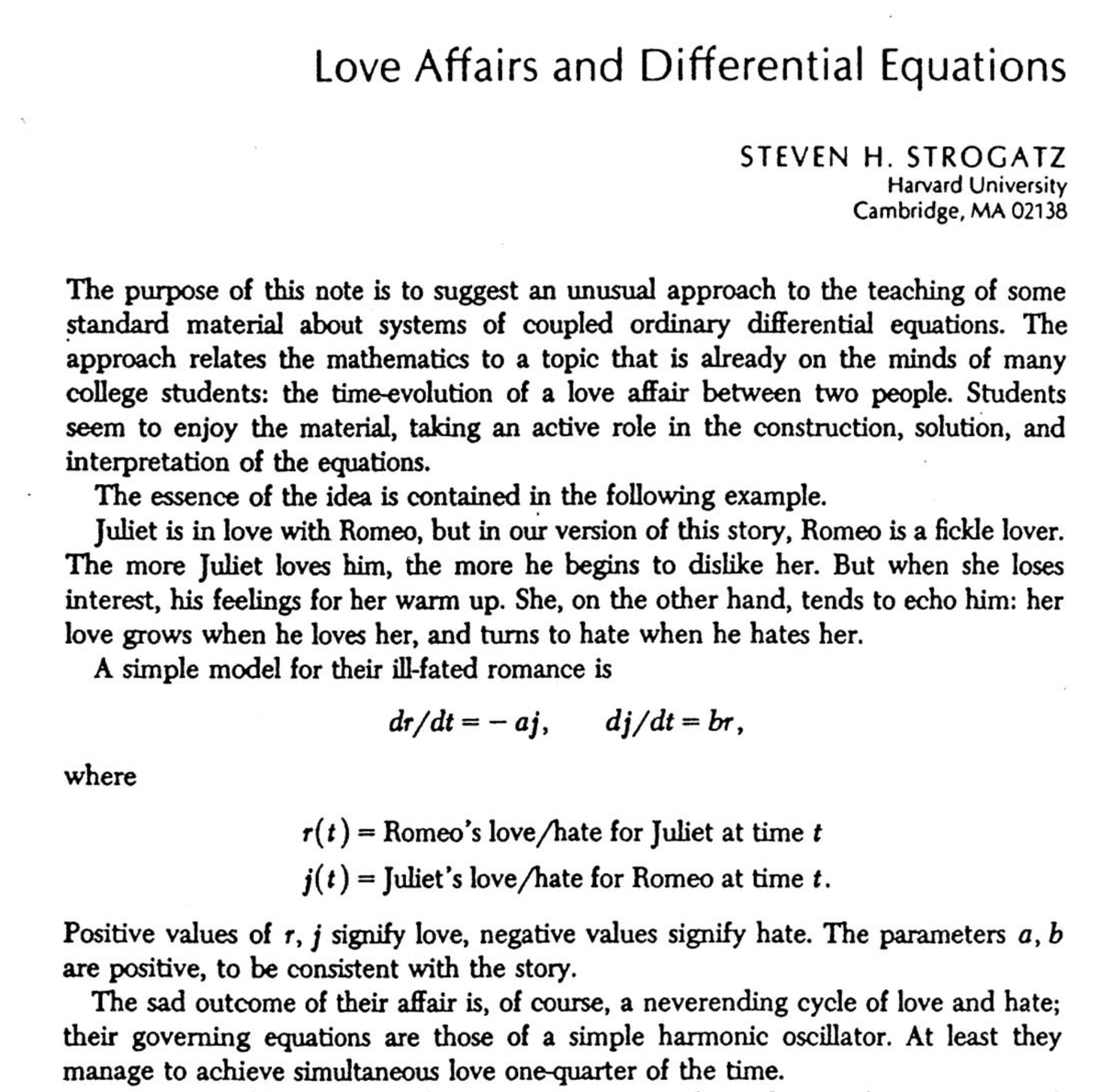











A list of papers that were rejected before going viral ( = winning a Nobel Prize). It just shows how #science works sometimes. ▫️ 1. Richard Ernst, Chemistry (1991), for NMR spectroscopy The paper that described our achievements was rejected twice by the Journal of Chemical Physics to be finally accepted and published in the Review of Scientific Instruments. ▫️ 2. Andre Geim, Physics (2010), for graphene “First, we submitted the manuscript to Nature. It was rejected and, when further information requested by referees was added, rejected again. According to one referee, our report did 'not constitute a sufficient scientific advance'." ▫️ 3. Paul Boyer, Chemistry (1997), for enzymatic mechanisms underlying the synthesis of ATP His proposed resolution of a major unsolved problem in biochemistry threatened to "change the paradigm," Boyer remembers, and "the leading journal" in his field - The Journal of Biological Chemistry - declined to publish his work. ▫️ 4. Herbert Kroemer, Physics (2000), for semiconductor heterostructures "I wrote up the idea and submitted the paper to Applied Physics Letters, where it was rejected. I was talked into not fighting the rejection, but to submit it to the Proceedings of the IEEE, where it was published, but ignored. I also wrote a patent, which is probably a better paper than the one in Proc. IEEE." ▫️ 5. John Polanyi, Chemistry (1997), for describing the dynamics of chemical elementary processes PRL rejected the paper as lacking scientific interest. Shortly thereafter they rejected T. Maiman's report of the first operating laser, on the same grounds. Polanyi read about this second rejection, quite by chance. [Later] he submitted the identical manuscript to the Journal of Chemical Physics, where it was promptly published. ▫️ 6. Kary Mullis, Chemistry (1997), for the PCR method (!!!!) "And Dan Koshland would be the editor of Science when my first PCR paper was rejected from that journal and also the editor when PCR was three years later proclaimed Molecule of the Year." ▫️ 7. Rosalind Yalow, Medicine (1977), for the radioimmunoassays of peptide hormones From the rejection letter: “The experts in this field have beer particularly emphatic in rejecting your positive statement that the "conclusion that the globulin responsible for insulin binding is an acquired antibody appears to be inescapable”. ▫️ 8. Hans Krebs, Medicine (1953), for the citric acid cycle The rejection letter from Nature is in the picture. 🤦♂️ ▫️ ❗ My point is simple: Rejections by editors are NOT rejections by the research community. Believe in your results. Bring them to the public. Post your study as a preprint. Show it to the world and let the world decide. ▫️ (This list compiled by Josh Nicholson + bit from me). #AcademicTwitter #phdlife