Ernesto J. Rojas
420 posts


Ernesto J. Rojas
@EJScience
Young Investigator @MaGIC_GCT | Chief Information Officer at @lmsa_west | @UCSF_MSTP ‘27 | @UCLA ‘18 | He/Him/Él | Views are my own | 🇭🇳






Professors have the best job in the world, especially when prior mentees like the phenomenal Ernesto Rojas come to visit. His career will be one to watch!







Registration for our LMSA West 2024 Regional Conference is now LIVE. If you’re a high schooler interested in medicine, premedical student or medical student you don’t want to miss out on this event! Buy tickets here: ucla.evenue.net/cgi-bin/ncomme…

Are you a medical or premedical student interested in presenting your research at our Regional Conference? Submit your abstract before our March 22 deadline! survey.zohopublic.com/zs/AFzEx3



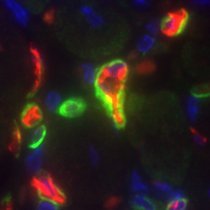


In scRNA-seq analysis, "overcorrection" of batch effects is a frequent cause for loss of important biological variability. Using prior celltype knowledge is a powerful way to limit overcorrection and should be more common practice. Sharing new preprint! biorxiv.org/content/10.110…