Dheeraj Prakaash me-retweet

excited to release a new benchmark for protein fitness prediction: FLIP2
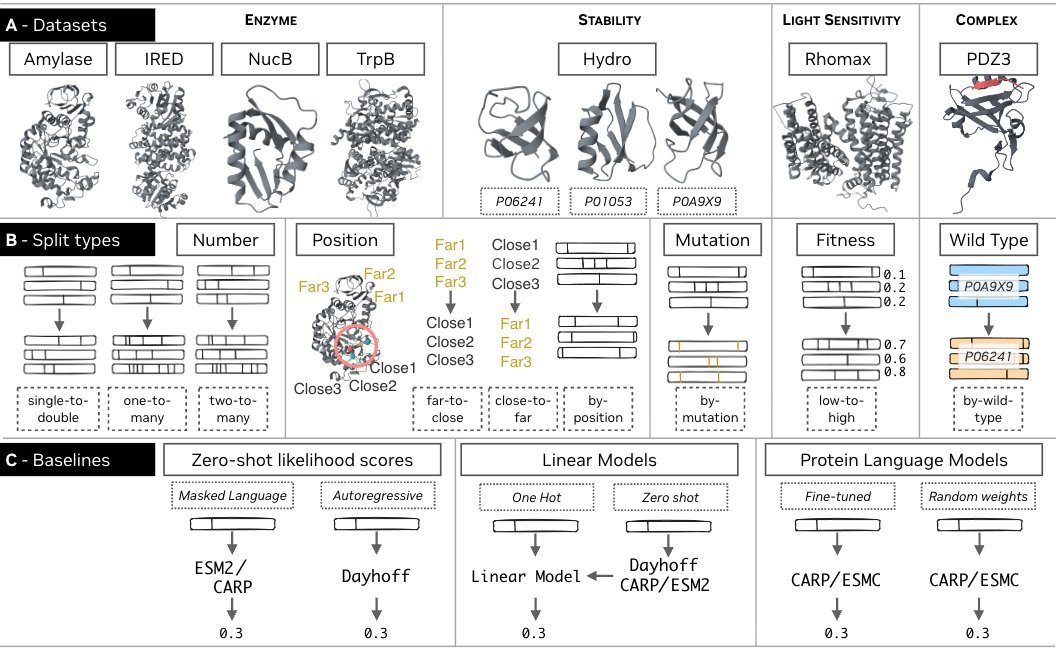
FLIP2 has 7 new datasets spanning enzymes, PPIs, and light-sensitive proteins, + splits designed to test generalization in realistic protein engineering settings
paper, data, code: flip.protein.properties
Kevin K. Yang 楊凱筌@KevinKaichuang
We made FLIP2, a protein fitness benchmark spanning seven new datasets, including enzymes, protein-protein interactions, and light-sensitive proteins, as well as splits that measure generalization relevant to real-world protein engineering campaigns.
English