Jules Samaran
33 posts

Jules Samaran
@JulesSamaran
PhD student @ Institut Pasteur. Research interests include optimal transport and computational biology.










The data science revolution is getting closer. TabPFN v2 is published in Nature: nature.com/articles/s4158… On tabular classification with up to 10k data points & 500 features, in 2.8s TabPFN on average outperforms all other methods, even when tuning them for up to 4 hours🧵1/19

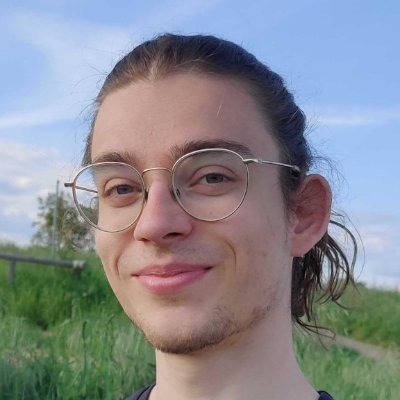
📽️On a interviewé @SibylleMarcotte , doctorante @ENS_ULM, membre de l'équipe Ockham, lauréate 🏆du prix Jeunes Talents France 2024 L'Oréal - @UNESCO #ForWomenInScience ▶️ses recherches et ses conseils pour les filles souhaitant devenir #scientifiques :) @UnivLyon1 @ENSdeLyon




Finally out a new paper from the lab lead by @TrimbourR "Molecular mechanisms reconstruction from single-cell multi-omics data with HuMMuS" doi.org/10.1093/bioinf… #singlecell #multilayernetworks

🥳 I’m very happy to announce our preprint biorxiv.org/content/10.110… ! scConfluence combines uncoupled autoencoders with Inverse Optimal Transport to integrate unpaired multimodal single-cell data in shared low dimensional latent space. @LauCan88 @gabrielpeyre
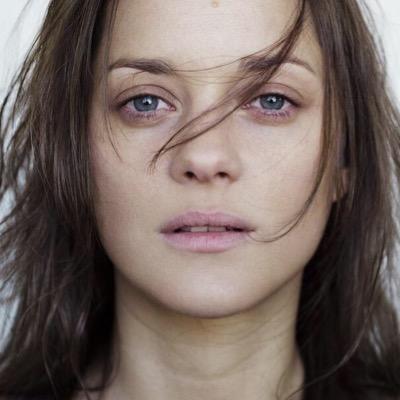













Newbies but goldies 😌: “Keep the Momentum: Conservation Laws beyond Euclidean Gradient Flows” with @gabrielpeyre and @RemiGribonval. We study conservation laws during the (euclidean or not) gradient or momentum flow of neural networks. arxiv.org/abs/2405.12888
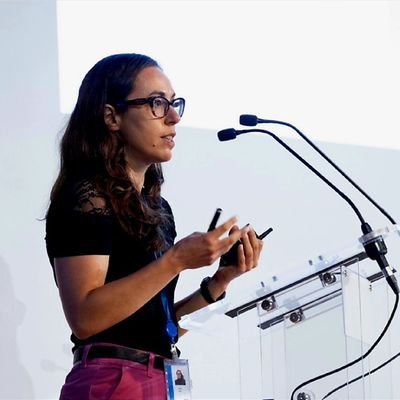





Looking for a postdoc to join my team @institutpasteur @CNRS @InstitutPrairie. The candidate will develop machine learning methods for single-cell omics data. Project funded by the #ERCStG MULTI-viewCELL


Finally out a new paper from the lab lead by @TrimbourR "Molecular mechanisms reconstruction from single-cell multi-omics data with HuMMuS" doi.org/10.1093/bioinf… #singlecell #multilayernetworks