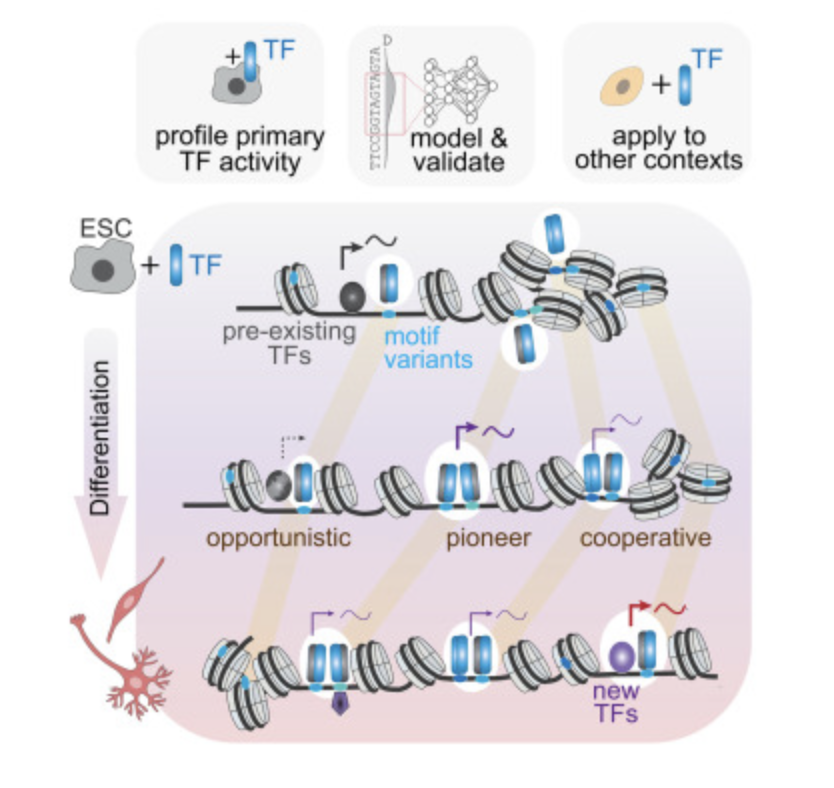
This year’s Ed Fischer Prize goes to Francesca Masoni (@SchubelerLab) for her PhD on chromatin organization—revealing how the ISWI chromatin remodeler subcomplex NURF positions nucleosomes to safeguard genomic boundaries & gene regulation. Read more at fmi.ch/news-events/ar…