Hugging Science
23 posts


Hugging Science
@huggingscience
The @huggingface community effort for science.


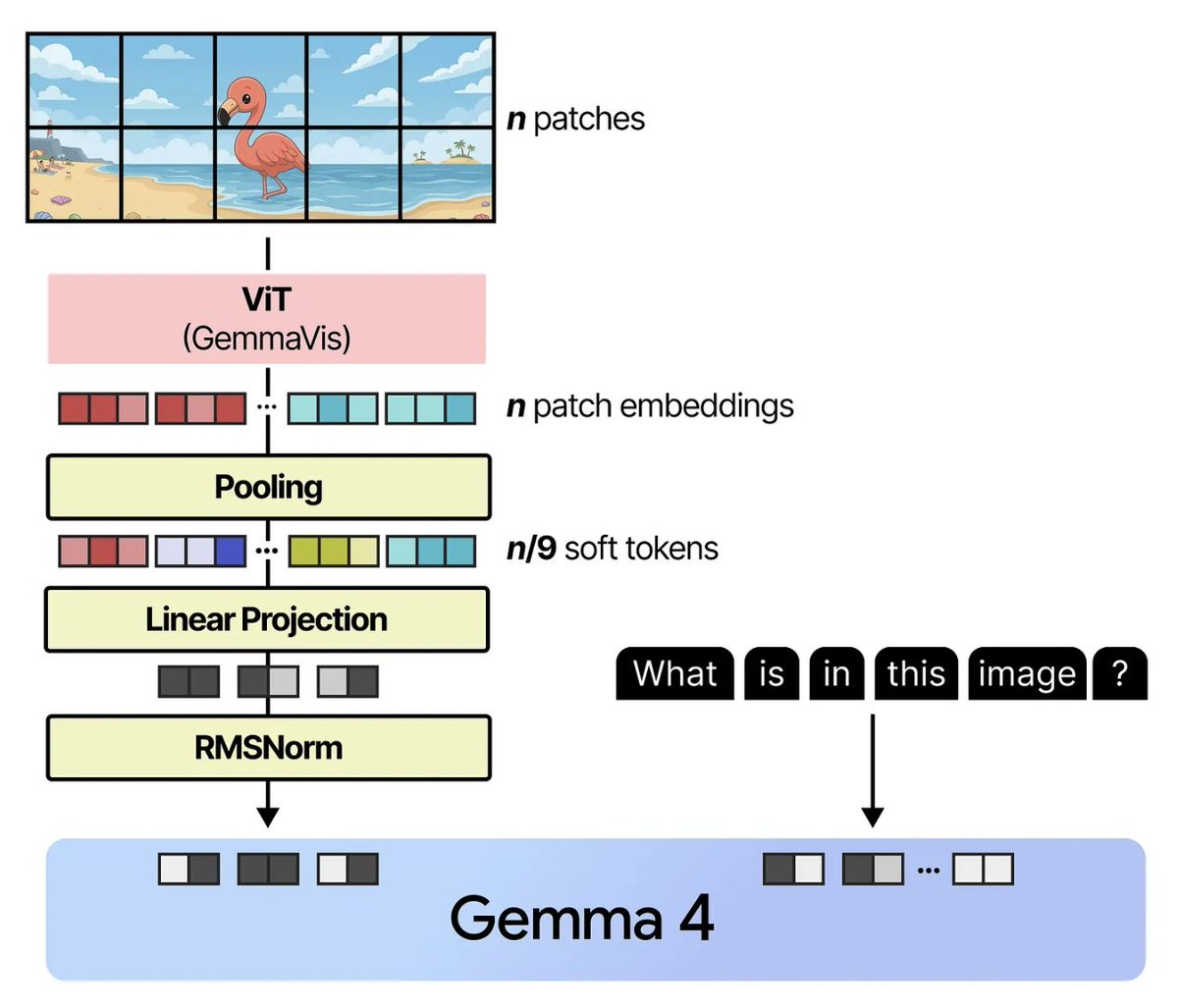
Most genomic AI models use fixed rules to process DNA into chunks, imposing arbitrary boundaries on a sequence with its own biological structure. @arnavshah0, @victor_ljz, and team developed dnaHNet, a tokenizer-free foundation model that learns its own segmentation from scratch, supervised by @_albertgu, @genophoria, and @BoWang87.

We’re excited to announce our collaboration with @huggingface. Through SAIR competitions, we aim to provide open data, benchmarks, tools, and models, and expand the frontier of AI x Science through collective contributions from the community. SAIR on Hugging Face: huggingface.co/SAIRfoundation











I’m co-organizing an “AI for Science: Algorithms to Atoms” social event during #NeurIPS2025 with Yann LeCun, Anima Anandkumar, Bill Dally, and Max Welling If you want to talk about AI Scientist, World Models and the future of AI-driven discovery, please come on Dec 5 3:30pm PT!

New preprint! We measured temperature- and pH-induced aggregation for over 18,000 natural and de novo designed protein domains!

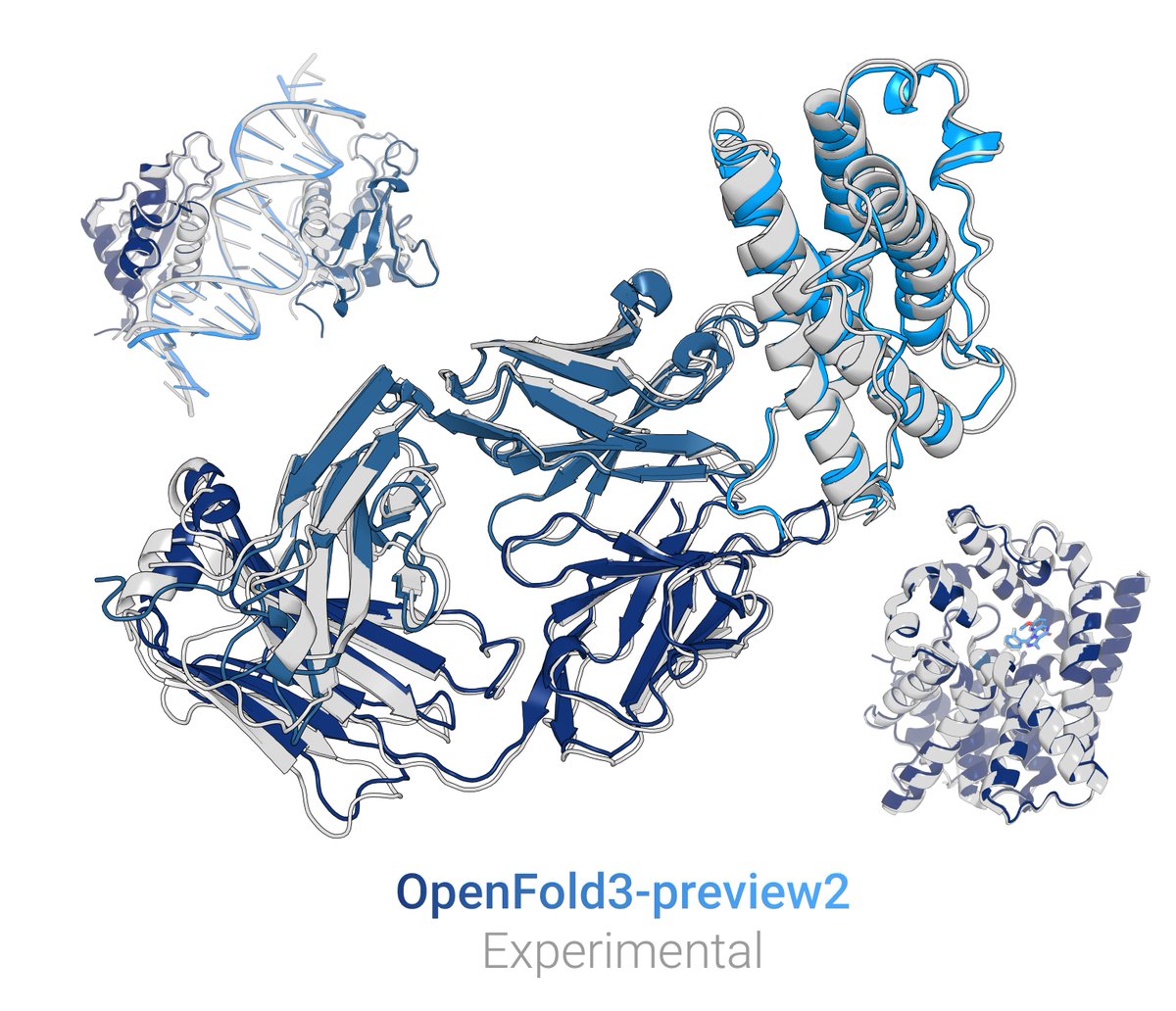
incredibly detailed technical blog just dropped on the anatomy of BoltzGen 🧬 made for ML people, but covering everything from molecular representations to diffusion-based generation of protein binders + crazy good interactive visuals 👏👏 @ludocomito