TF binding papers 📚 NucPosDB database 리트윗함
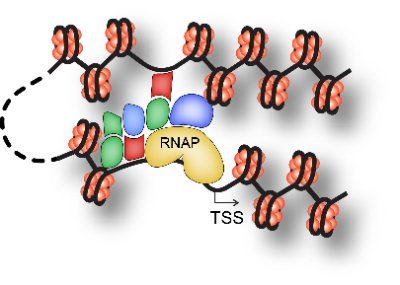
Footprint-C reveals transcription factor modes in local clusters and long-range chromatin interactions [Liu et al, 2024] nature.com/articles/s4146…
▶️At one-dimensional level - modes of TF co-occupancy
▶️At pairwise contact level - multiway chromatin interactions of different TFs




English