Sabitlenmiş Tweet

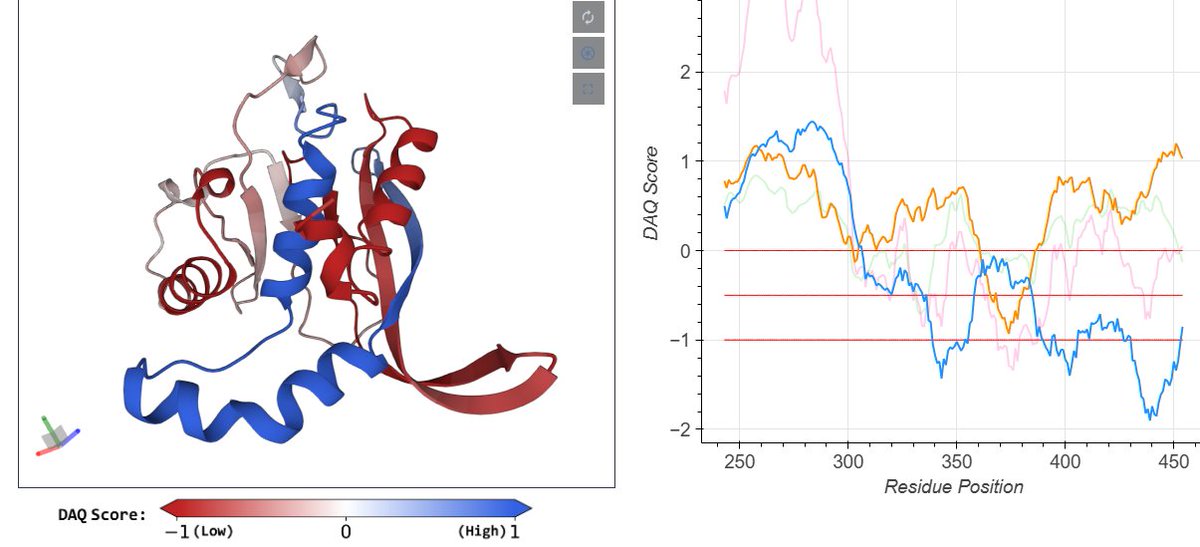
Very excited to share our new paper DAQ-score, just published in @naturemethods .
Our deep learning method can estimate the local quality of the protein structure model from cryo-EM data.
DAQ is available on Google Colab:
bit.ly/daq-score
Paper:
lnkd.in/eTvJfSvH
English