Karsten Rippe
164 posts



9/ 🤘 It's a long way to the top if you wanna rock 'n' roll... but @IsabelleSeufert rocked it with help of great PhD/Master's students @DKFZ & BioQuant, and valuable contributions from @AkisPapantonis, @jpmallm1, @s_anders_m & Petros Kolovos labs & funding by @spp2202_dfg. #ACDC
English

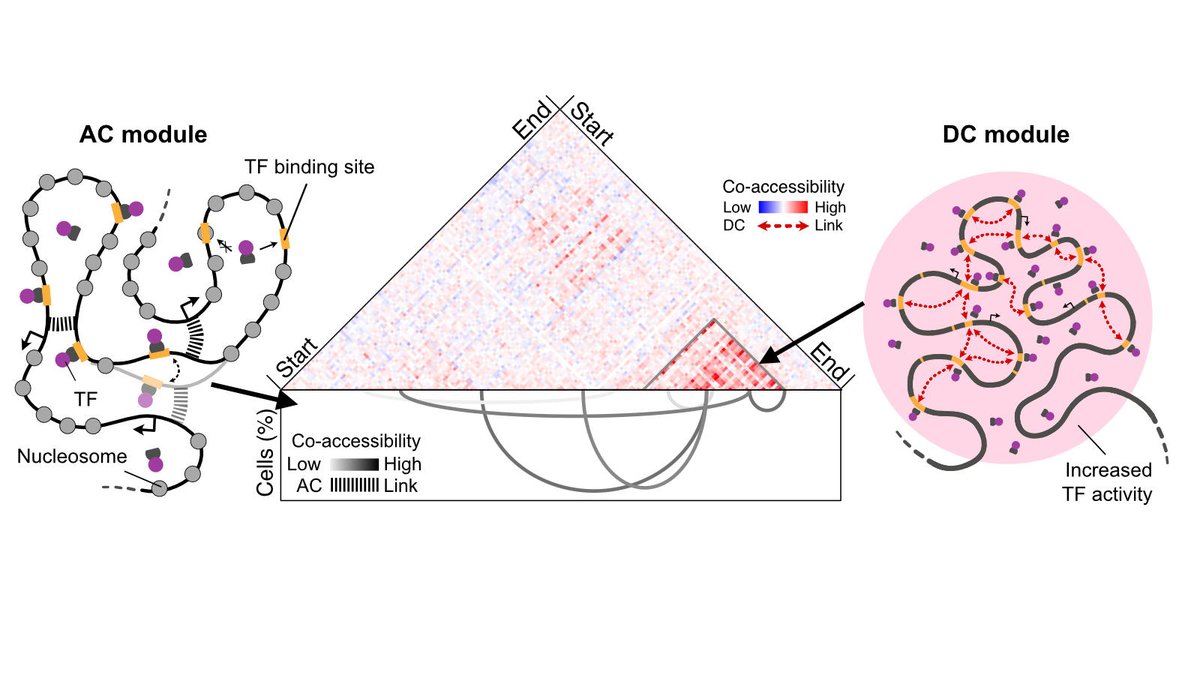
8/ 🔥 We reconcile findings from previous sequencing and microscopy studies into the genome-wide AC/DC model of #transcription regulation, revealing two co-existing chromatin compartment architectures. For the full story, check out our bioRxiv preprint at doi.org/10.1101/2024.0….
English

1/ 🤔 Do #chromatinlooping-mediated promoter-#enhancer interactions drive gene expression? Or is the local enrichment of transcription factors into hubs/#condensates the key factor? Our AC/DC model of gene regulation at bioRxiv doi.org/10.1101/2024.0… provides a new view on this.

English

Great joint effort with @jpmallm1 from the Single Cell Open Lab @DKFZ and driven by Anne Rademacher, @alik_huseyn0v and Michele Bortolomeazzi with valuable contributions from our collaborators @KiTZ_HD and crucial funding from the MULTI-SPACE program of @ResearchLifesci.
English

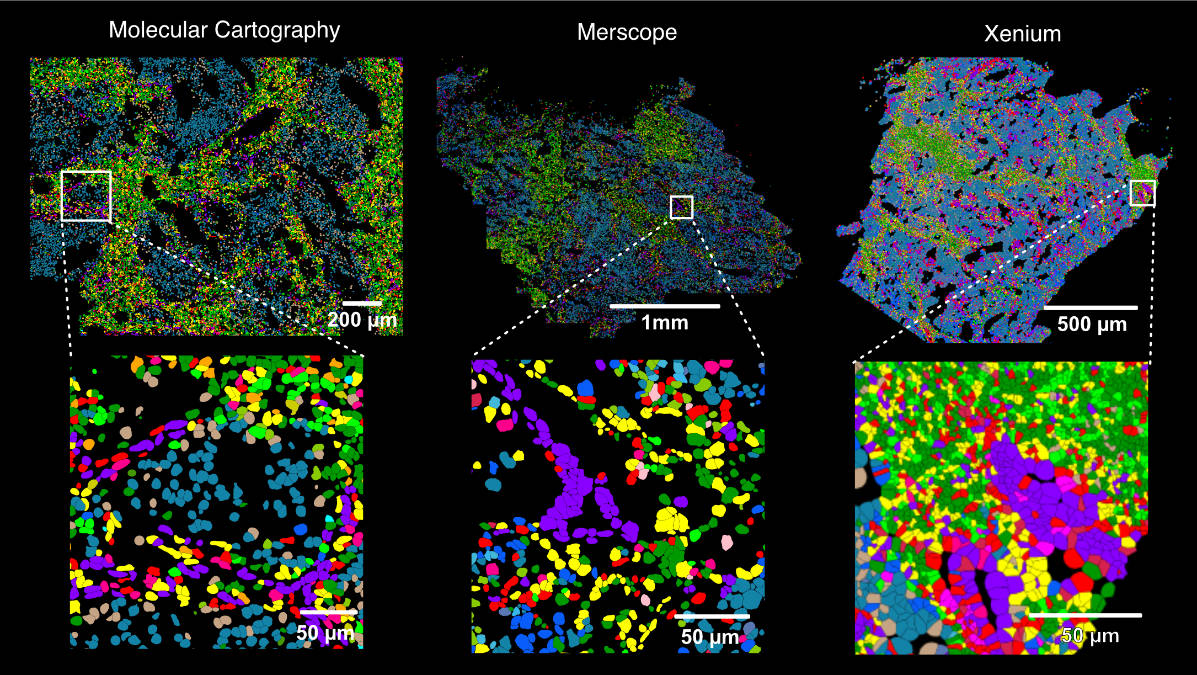
Which of the different spatial transcriptomics (ST) technologies should you use for your research? The lessons we have learnt from comparing RNAscope, Molecular Cartography (MC), Merscope, Xenium and Visium are now on bioRxiv doi.org/10.1101/2024.0… 🧵
English
Karsten Rippe ретвитнул

Registration is open! Join us in October for our quadrennial international #chromatin symposium in Munich. Hear and meet amazing guest speakers, and present your own work. Details at sfb1064.med.uni-muenchen.de/symposium2024/…

English
Karsten Rippe ретвитнул

Do you want to do excellent #postdoc research supported by one of the most prestigious fellowships in 🇪🇺? Do you want to work at a leading institution with world-class scientists from all over the world? Then apply to the DKFZ #MarieCurie MasterClasses via t1p.de/l01hq
English

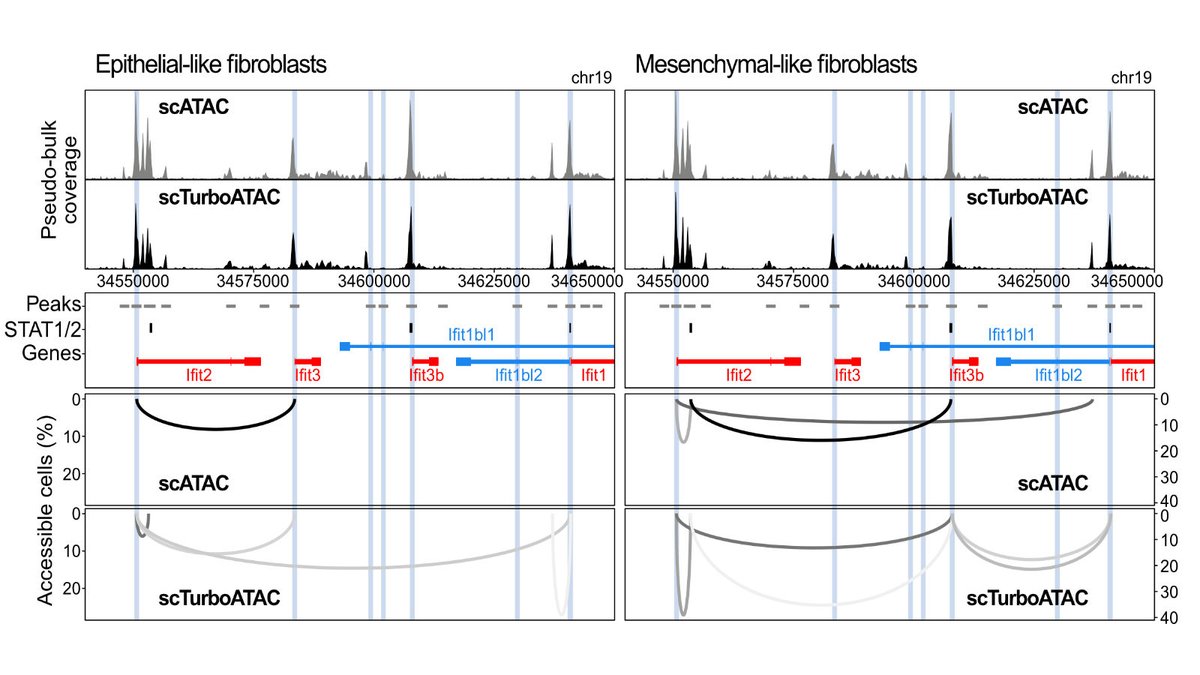
Enjoying the work with @IsabelleSeufert and @jpmallm1 to increase the coverage of scATAC-seq for co-accessibility analysis and other applications frontiersin.org/articles/10.33…. Our latest data are at 150k unique fragments/cell. Stay tuned...
English

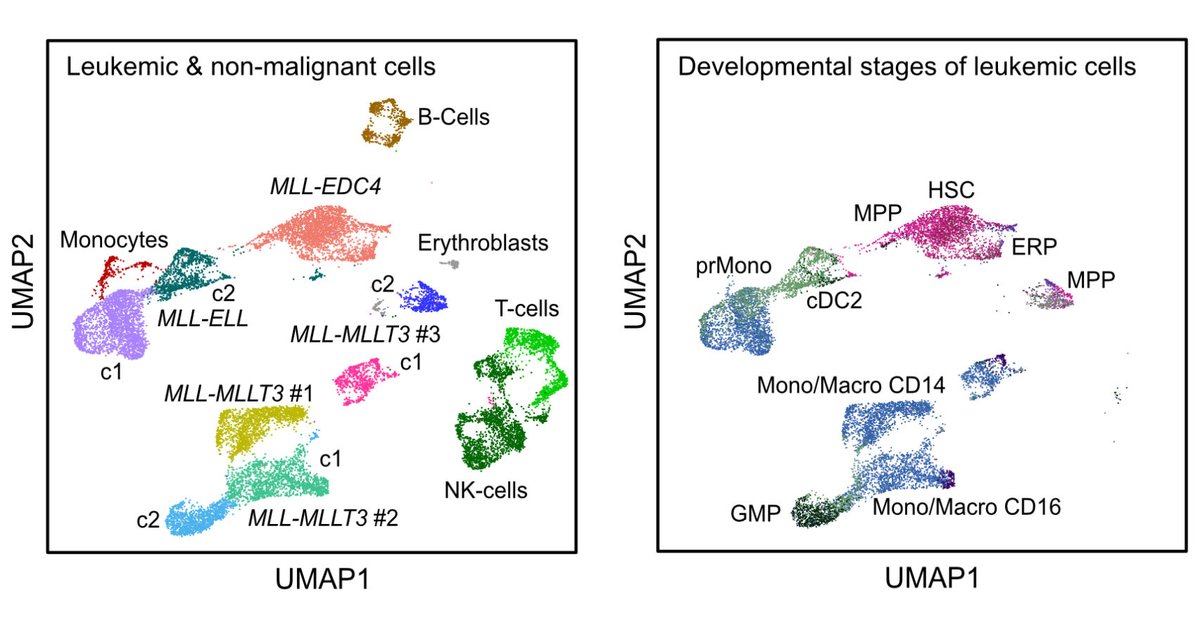
Our single cell analysis of MLL/KMT2A fusions in AML with Michael Lübbert's group @Uniklinik_Fr is now out in @BloodAdvances. It shows a progenitor-like phenotype of the novel MLL-EDC4 fusion. Free download link: authors.elsevier.com/sd/article/S24…. Thanks to DFG FOR2674 for support.
English
Karsten Rippe ретвитнул

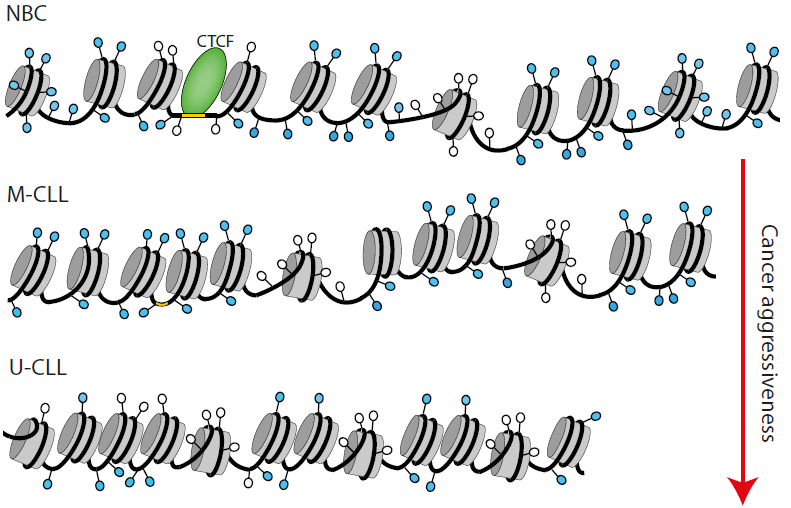
Our paper on nucleosome repositioning in CLL is now published in @genomeresearch! Led by @krissyvalp in our lab, collab with @KostareliLab, @KarstenRippe genome.cshlp.org/content/33/10/…
▶️Aggressive cancers have shorter nucleosome spacing
▶️Distinguish cancer subtypes from nuc patterns
English
