Varosync
5 posts


@var0sync
Varosync is a computational drug development company building AI systems to model complex biology and optimize therapeutic design.





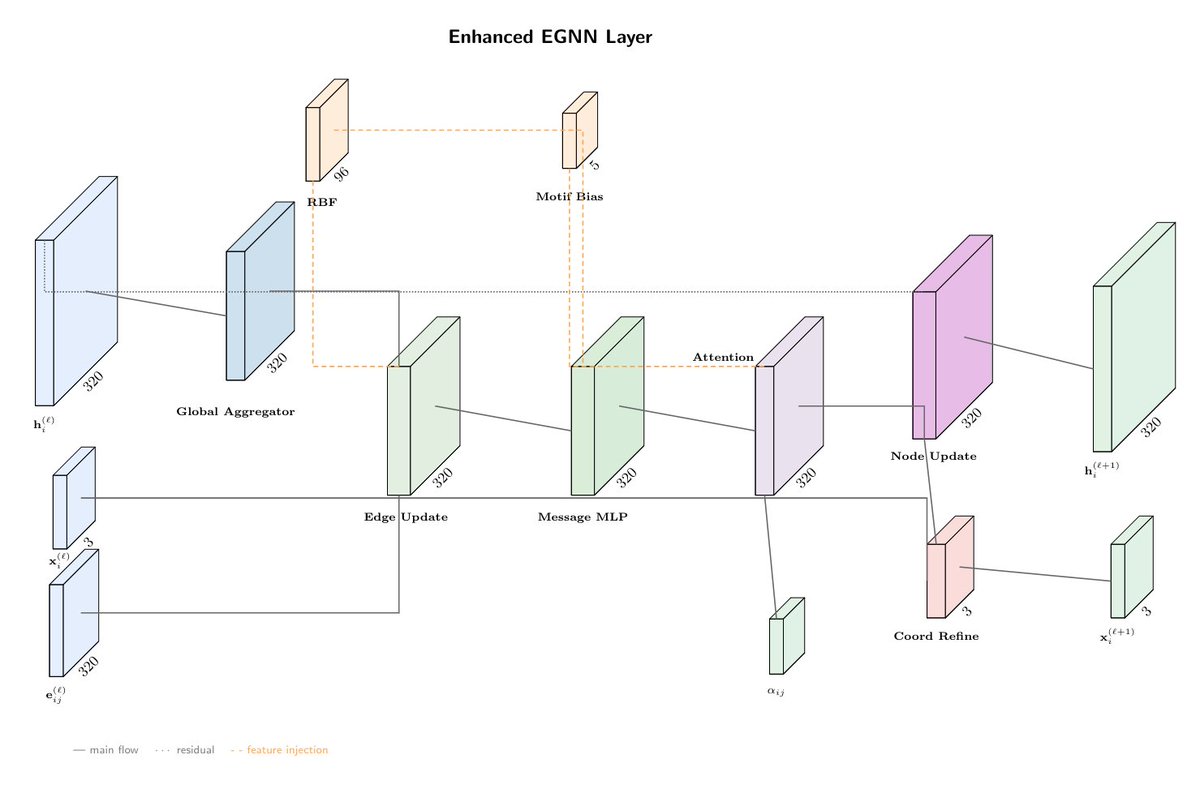
Hyaline: Geometric Deep Learning for Accurate Prediction of G Protein-Coupled Receptor Activation States from Structure 1. Hyaline introduces a novel geometric deep learning framework that predicts GPCR activation states with near-perfect accuracy (AuROC 0.995) by integrating E(n)-equivariant graph neural networks and ESM3 evolutionary embeddings. This approach bridges the gap between sequence-based and structure-based methods, outperforming sequence-only models significantly. 2. The model leverages E(n)-equivariant message passing to capture the subtle, non-local conformational changes that distinguish GPCR activation states. This ensures that predictions are invariant to rotations and translations, focusing solely on the relative arrangement of atoms in the receptor structure. 3. Hyaline incorporates biological priors through motif-specific attention biasing, prioritizing residues within conserved activation motifs such as the DRY, NPxxY, and CWxP motifs. This not only accelerates learning but also improves robustness and interpretability, with attention weights revealing key regions involved in receptor activation. 4. The model demonstrates robust generalization across diverse GPCR classes, including challenging Class C receptors, and maintains high performance even on structures determined with the latest cryo-EM methods. This versatility makes Hyaline a powerful tool for high-throughput discovery of allosteric modulators. 5. Ablation studies highlight the importance of each architectural component, confirming that both evolutionary embeddings and geometric message passing are essential for accurate state discrimination. The model’s attention mechanisms provide insights into the structural basis of activation, correlating strongly with known activation signatures. 6. Error analysis reveals that most misclassifications reflect genuine biological ambiguity rather than model failure, with intermediate states representing opportunities for developing functionally selective compounds. Hyaline’s ability to identify these states could accelerate the discovery of conformationally selective drugs. 7. Hyaline’s linear computational scaling and efficient implementation enable rapid inference on typical GPCR structures, facilitating high-throughput screening of AI-predicted structures. This capability is crucial for assessing the activation states of computationally generated GPCR models in drug discovery contexts. 📜Paper: biorxiv.org/content/10.648… #GPCR #GeometricDeepLearning #ProteinStructure #ActivationStatePrediction #DrugDiscovery #AIinBiology