Behnam Yousefi ری ٹویٹ کیا

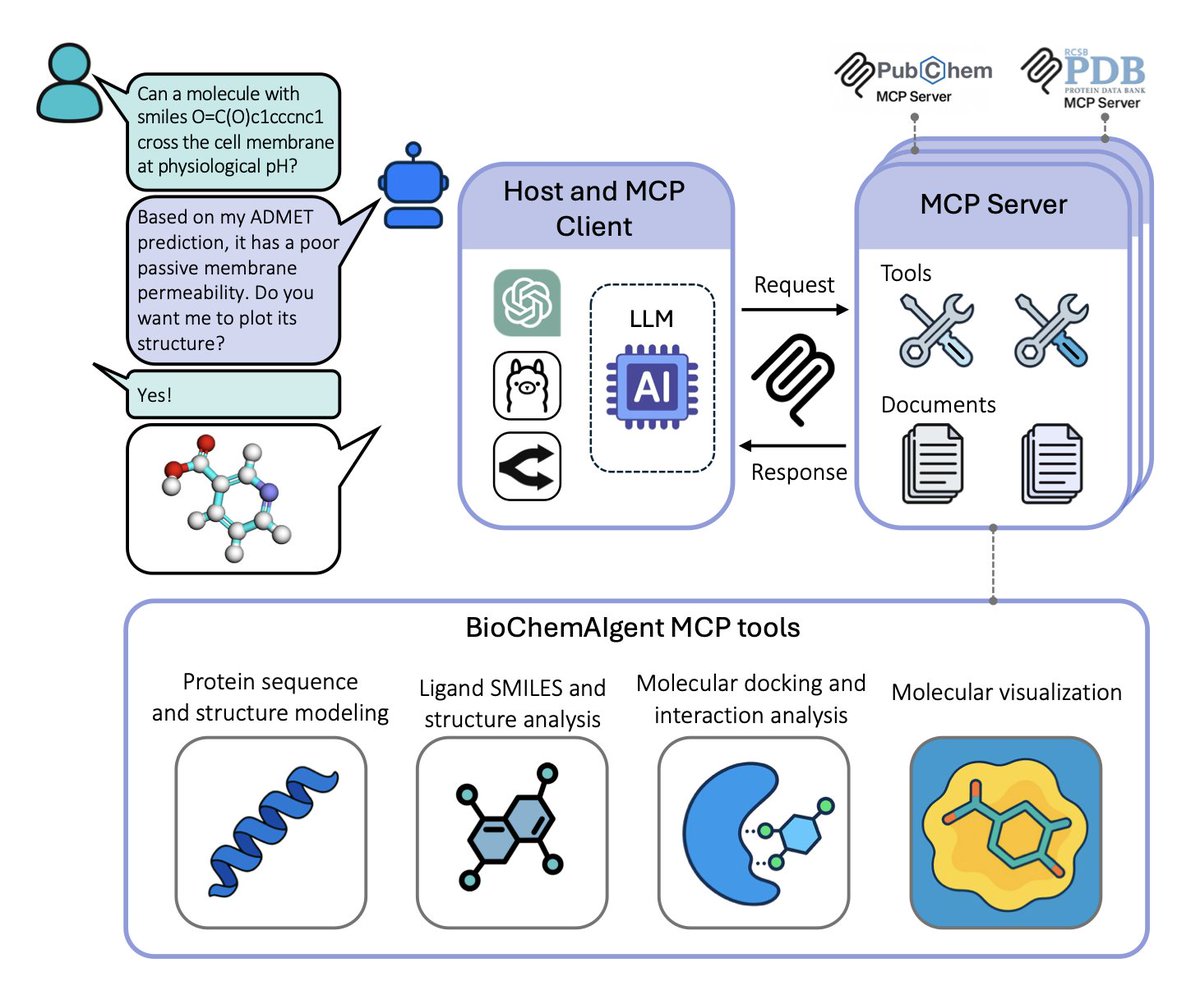
BioChemAIgent: An AI-driven Protein Modeling and Docking Framework for Structure-Based Drug Discovery
1. First agentic framework that unifies 19 tools—ESM3, AlphaFold3, AutoDock Vina, DiffDock, etc.—into one chat-style interface for end-to-end small-molecule discovery, slashing the usual multi-software integration burden.
2. Wraps complex pipelines (protein prep, protonation, grid setup, docking, interaction fingerprinting, 3-D visualization) into reproducible, transparent workflows that non-coders can launch with plain English.
3. Built-in “render_structures” and “interaction_plot” modules translate natural-language style requests into publication-grade 3-D images, eliminating the need for PyMOL scripting expertise.
4. Dual evaluation strategy: 98.5 % automatic accuracy on 65 corrupted queries plus perfect expert scores on four real-world case studies, showing GPT-5-powered agent rivals trained chemists in task planning and result interpretation.
5. Community-oriented registry on GitHub invites developers to plug in new MCP servers, turning the system into a living, expandable ecosystem rather than a frozen black box.
6. Open web interface already live—users can dock ibuprofen to COX-1, visualize the Arg120 salt bridge, and download inputs/outputs without installing anything.
💻Code: github.com/imsb-uke/bcai
📜Paper: biorxiv.org/content/10.648…
#DrugDiscovery #AI #StructuralBiology #MolecularDocking #AlphaFold3 #ESM3 #ChemInformatics #OpenScience
English