RGB Lab @ MIT
151 posts


RGB Lab @ MIT
@RGBLabMIT
Official Twitter for the Gómez-Bombarelli group @MIT_DMSE | We use atomistic simulations and ML for accelerated materials design | Managed by group members



This paper shows that wildly different AI models for molecules, materials, and proteins are independently learning the same underlying representation of matter suggesting we’re converging on a shared physics-grounded “latent reality” that proves scientific foundation models might actually be generalizable across domains


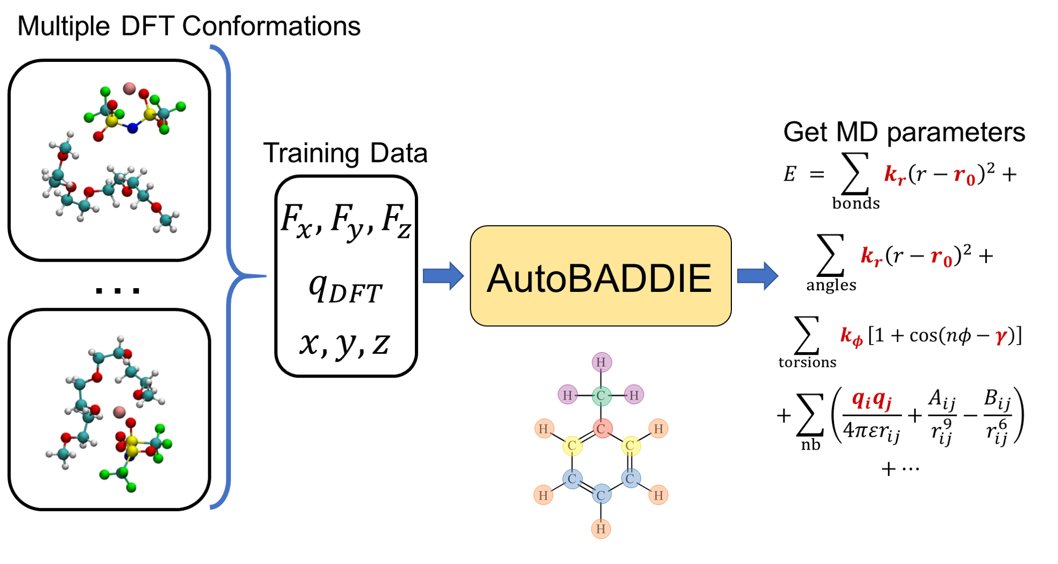
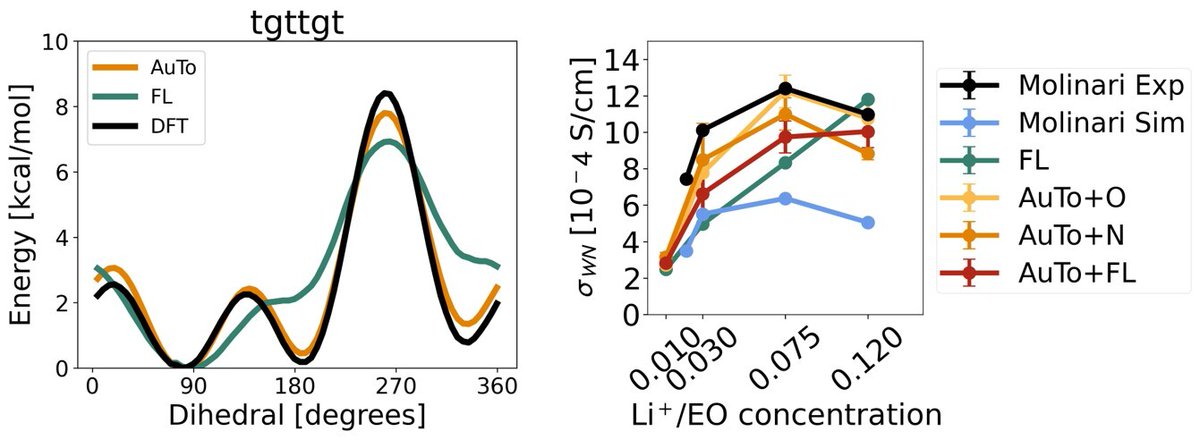


The surprising ineffectiveness of molecular dynamics coordinates for predicting bioactivity with machine learning 1. The study challenges the assumption that molecular dynamics (MD)-derived coordinates are superior for machine learning-based bioactivity predictions, revealing that they often underperform compared to minimum-energy conformations. 2. Using over 2600 protein-ligand complexes, the authors systematically compared MD-derived and minimum-energy coordinates, employing three descriptor sets and machine learning algorithms like Random Forest, XGBoost, and Support Vector Regression. 3. Surprisingly, MD-derived conformations failed to consistently outperform minimum-energy structures, even though they provide dynamic representations of molecular interactions. 4. In certain cases, ensemble averaging of MD-generated snapshots improved predictive performance slightly, but the benefits were not proportional to the computational costs. 5. Extended Connectivity Fingerprints (ECFPs), a 2D molecular representation, outperformed MD-based models in many cases, questioning the utility of complex 3D data for predicting bioactivity. 6. The findings highlight a critical need for better 3D and dynamic molecular representations. The study suggests exploring geometric deep learning or incorporating protein information to improve machine learning models. 7. The authors propose a tiered approach: using fast, simpler methods like ECFPs for initial screening, followed by MD-based predictions for refining top candidates. 8. The study serves as a wake-up call for molecular machine learning, emphasizing the importance of balancing data complexity, computational cost, and predictive accuracy. @fra_grisoni @DerekvTilborg @Rza_ozcelik 📜Paper: chemrxiv.org/engage/chemrxi… #MachineLearning #DrugDiscovery #MolecularDynamics #Bioinformatics #AI





Instead of the typical (somewhat technical) lecture, we discussed a couple of papers, including a favorite J. Chem. Ed. activity of mine: "Chemistry from Telephone Numbers: The False Isokinetic Relationship." It's a lot of fun. Do check it out. 2/3 pubs.acs.org/doi/abs/10.102…









