ShenoyLab
215 posts


ShenoyLab
@ShenoyLab
We develop theoretical concepts and computational models - mechanobiology, matrix mechanics, nano- and energy- materials. Tweets by group members. @CEMB_STC
University of Pennsylvania انضم Haziran 2020
197 يتبع1.1K المتابعون

6/6 🤝 Huge thanks to @ShenoyLab, @Melike_Lak, and @rp_mccord for the collaboration, and @CEMB_STC, the NCI MetNet - @NCIDCB! This work highlights the power of interdisciplinary science.
English

1/6 We built a sequencing-informed polymer model that integrates STORM, Hi-C, and RNA-seq to reveal how HC domains — especially their boundaries —regulate gene expression during drug treatment, microenvironmental changes, and cell fate transitions. doi.org/10.1038/s41467…

English
ShenoyLab أُعيد تغريده

ShenoyLab@ShenoyLab
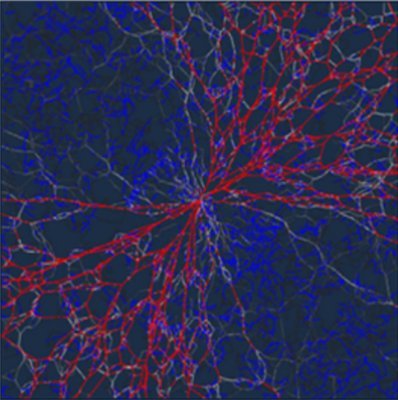
A new joint theory/experiment paper led by @ZeGongGreg just appeared in Nature Communications! We show how wave-like dynamics of immune cell podosomes arise from the coupling between chemo-mechanical signaling and actin diffusion. @CEMB_STC @PennEngineers nature.com/articles/s4146…
QHT
ShenoyLab أُعيد تغريده

Great news: the next meeting of the Invadosome Consortium (9th in this series) will take place in Vancouver, Canada, Oct. 01-04, 2024
Organizers: Karla Williams and Michael Olson
Excellent science and beautiful nature, what´s not to like? ⚗️🌲
invadosome.com

English
ShenoyLab أُعيد تغريده

Incredible speaker lineup uncludes @KoleRoybal, @NF_Steinmetz, @KWhiteheadLab, @NatalieArtzi, @LindaGGriffith1, @ManishButte, @Tang_Lab_UCSF, @ValerieJHorsley , @BellasFATLab, @taabaman, @ShenoyLab, @DoloffLab, @SantiCorreaPhD, @JennyNJiang, @gail_l_rosen, @veiseho and many more!
English
ShenoyLab أُعيد تغريده

Come one, come all, I want to see you here this Fall! Register for #IMES2024!
Trainees & early career investigators - submit your abstracts for a chance to win cash prizes and oral presentations! Travel Scholarships are available!🤑
Due: Oct. 1
Link: tinyurl.com/4swvjttr
Drexel School of Biomedical Engineering@drexelbiomed
🚨 Last Call: Abstract Submission Deadline Approaching! 🚨 #IMES2024 is the only conference dedicated to convergent research in translational immunology and engineering. 📅 Deadline: Oct. 1, 2024 Submit here: tinyurl.com/35uwzcj7 Learn more: tinyurl.com/IMES2024
English
ShenoyLab أُعيد تغريده
ShenoyLab أُعيد تغريده
What balances the size of heterochromatic domains in our cells? @ShenoyLab et al explore the interplay of methylated histones, acetylation, and RNA polymerase II-driven supercoiling. Check out the sophisticated model led by Aayush Kant on @NatureComms nature.com/articles/s4146…

English
ShenoyLab أُعيد تغريده

Last day of the wonderful GRC on Signal Transduction by Engineered Extracellular Matrices #GRCSTEEM organized by Shelly Peyton and @kumarlabucb started with a session on multiscale modelling, with discussion leader Vivek Shenoy and first speaker @davidodde


English

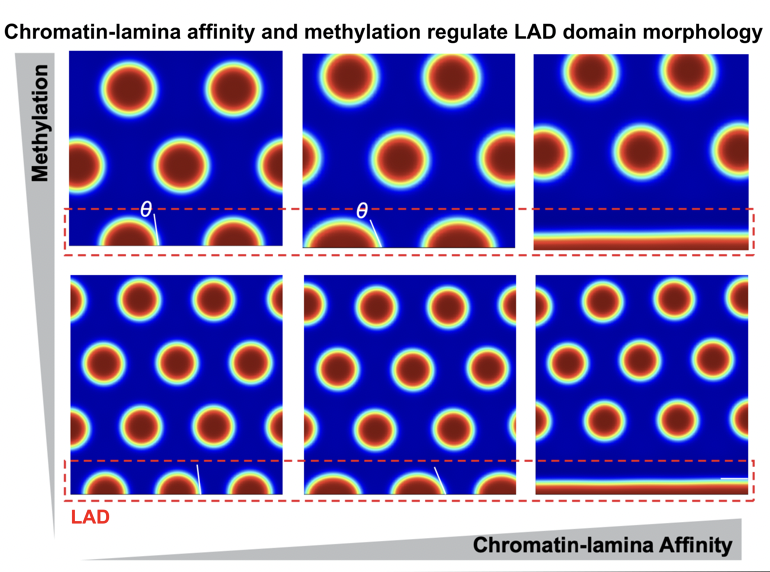
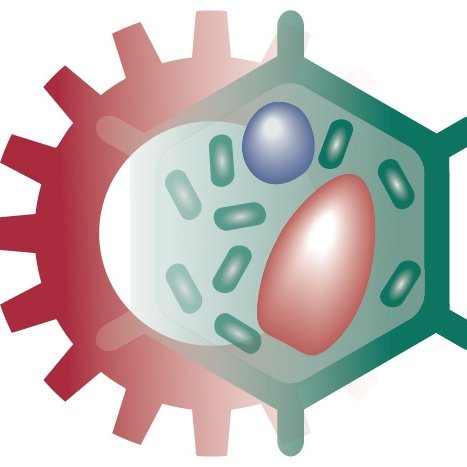
@Ella_Maru Thank you! Yes, that is what we show here. Substrate stiffness changes HDAC3 levels, which in turn alters the chromatin-lamina interaction strengths. Note HDAC3 is involved in "tethering" chromatin to the lamina.
English

@ShenoyLab Impressive study! Could differences in LAD morphology be due to changes in chromatin-lamina interaction strength across substrate stiffnesses?
English

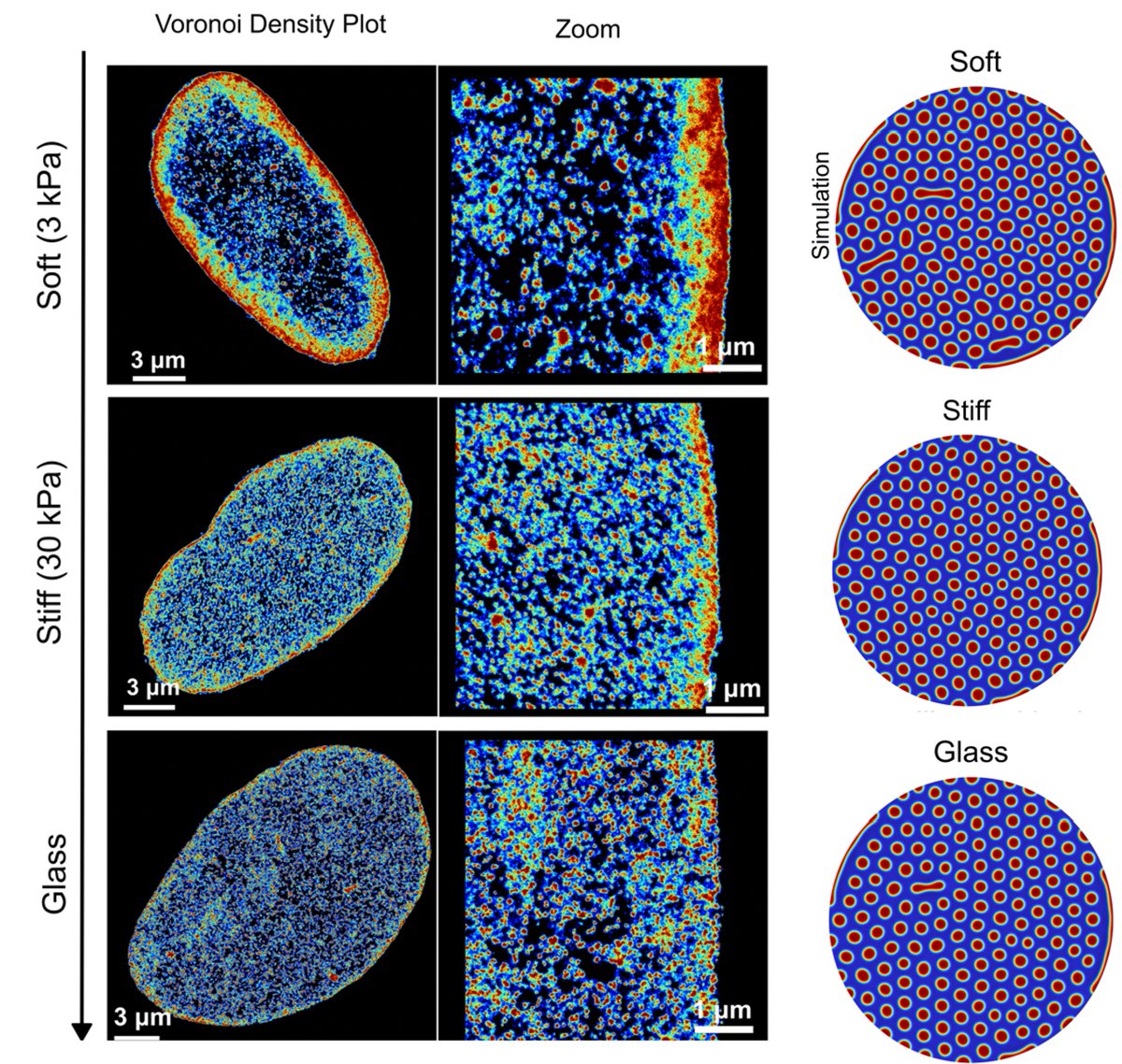
New preprint on integrating biophysical computational modeling with high-resolution imaging to predict how epigenetics and chromatin-lamina interaction energetics regulate the size and shape of lamina-associated domains (LADs)! biorxiv.org/content/10.110…
English

By observing the size and shape of LADs via high-resolution STORM imaging by our amazing collaborators @Melike_Lak @HeoLab and @MauckLab, we can quantitatively extract the biophysical parameters - epigenetic reaction rate and chromatin-lamina affinity.
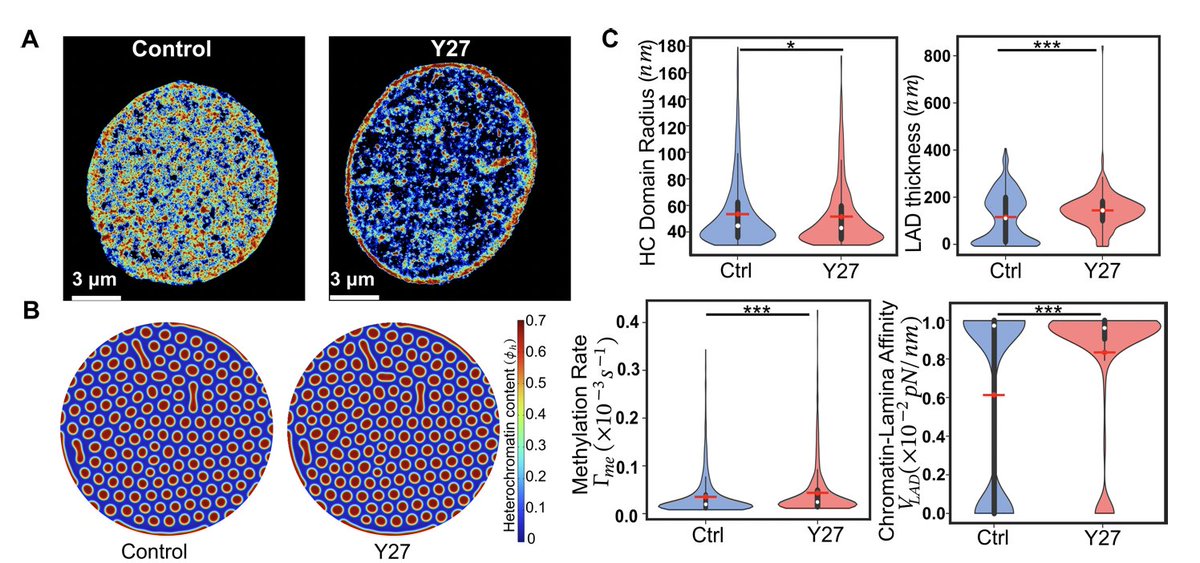
English