Pierce Lab أُعيد تغريده

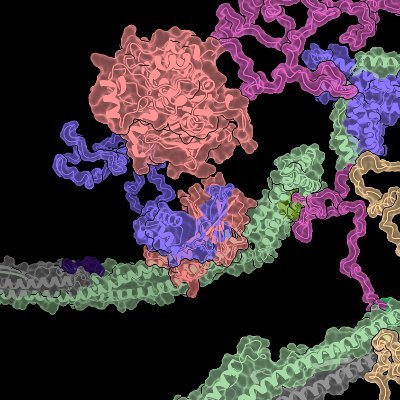
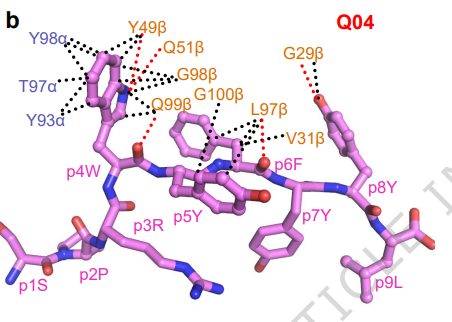
Structural insights into clonal restriction and diversity in T cell recognition of two immunodominant SARS-CoV-2 nucleocapsid epitopes @NatureComms 🇺🇸🇨🇳
nature.com/articles/s4146…
English
Pierce Lab
130 posts

@pierce_lab
Computational Structural Biology, Immune Recognition, Protein Design @UMD_IBBR and University of Maryland CBMG Dept







BREAKING NEWS The Royal Swedish Academy of Sciences has decided to award the 2024 #NobelPrize in Chemistry with one half to David Baker “for computational protein design” and the other half jointly to Demis Hassabis and John M. Jumper “for protein structure prediction.”