Robert King
1.3K posts

@rob234king
Experienced Senior Bioinformatician primarily working with sequencing data but can solve most tasks if the data is good. Opinions are all my own.




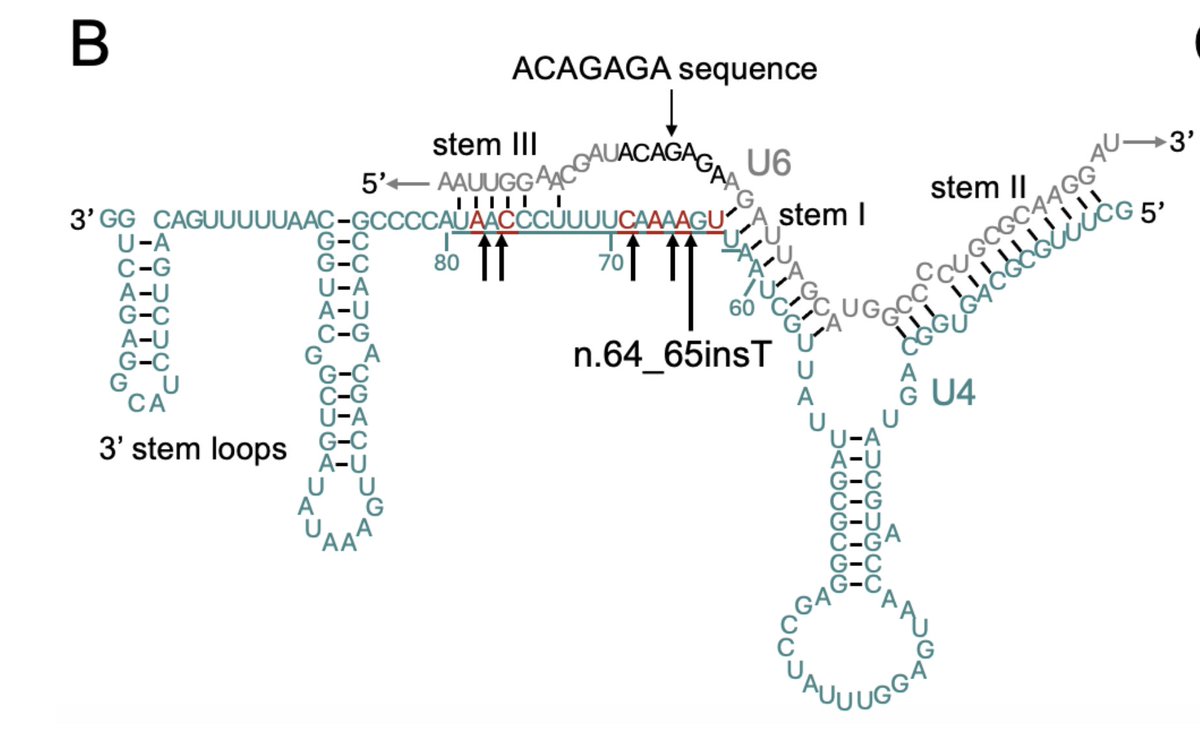
Would you believe me if I told you that a single variant in a non-coding RNA explains ~0.5% of all undiagnosed individuals with neurodevelopmental disorders (NDD) in @GenomicsEngland ??? I didn’t initially either, but here is the story of RNU4-2 🧵1/9

We've got plenty to look forward to at #AACR24. Drop by booth 3053 throughout the event to meet our team of experts. Be sure to join our Spotlight Session & live demos hear more about how nanopore sequencing is accelerating oncology research. Learn more: bit.ly/3TFVvvj




Minimap2-2.27 released with new presets for accurate Nanopore reads and Illumina Complete Long Reads (both recommended by vendors), along with a few minor features and bug fixes. Output alignment identical to v2.26 except the value at a custom tag. github.com/lh3/minimap2/r…







Our preprint “High resolution long-read telomere sequencing reveals dynamic mechanisms in aging and cancer” is now on bioRxiv. biorxiv.org/content/10.110…