
Alan DenAdel
335 posts

@AlanDenadel
Bioinformatics scientist @AllenInstitute. PhD in computational biology from @BrownCCMB @BrownUniversity. Previously at @illumina. he/him









New preprint: Beyond alignment: synergistic integration is required for multimodal cell foundation models. 🔗 biorxiv.org/content/10.648… Multimodal compositional cell foundation models are emerging as a path toward virtual cells. But when does multimodal fusion truly add info? 🧵






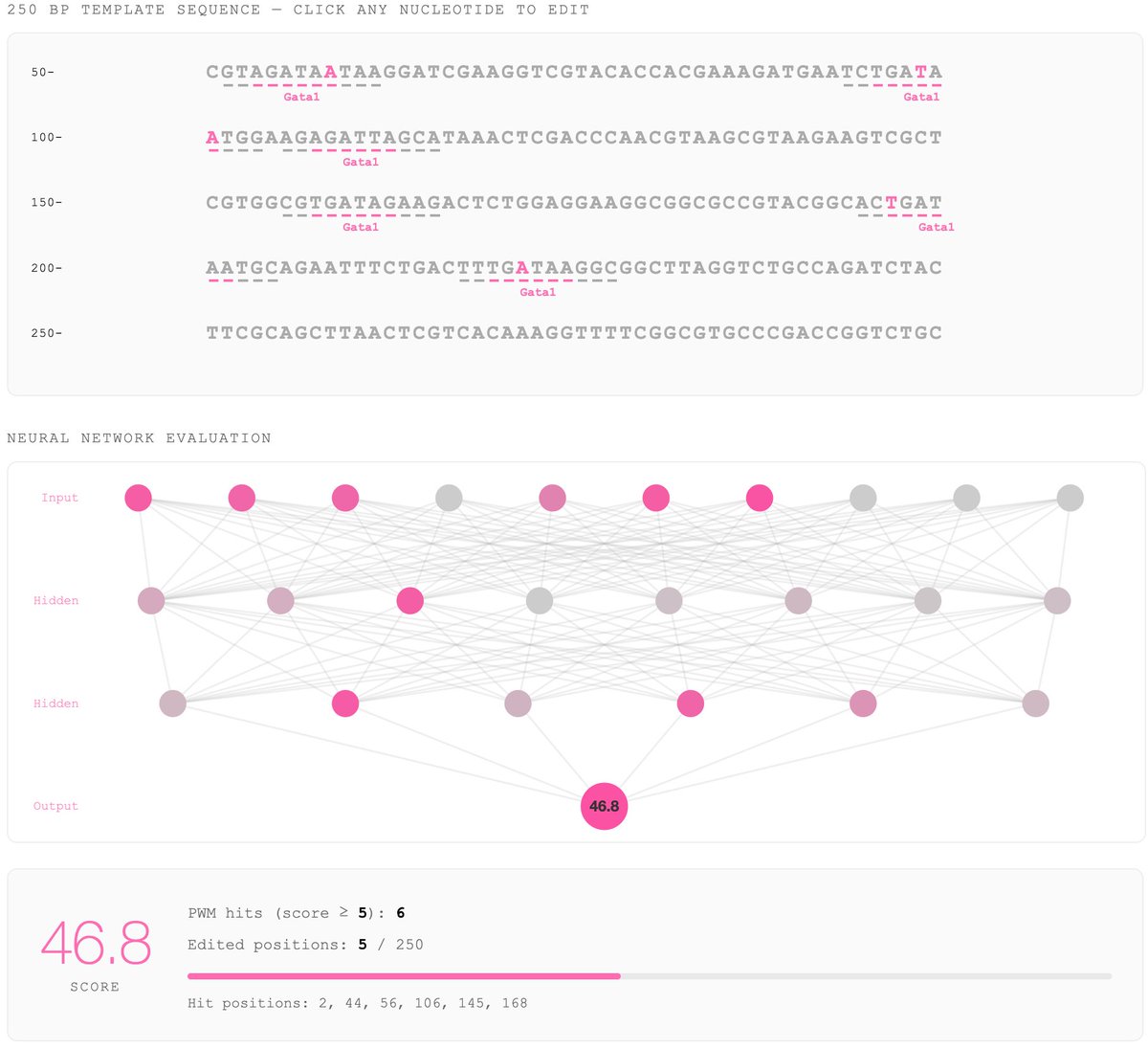
Announcing our preprint: Predicting evolutionary rate as a pretraining task improves genome language model representations: biorxiv.org/content/10.648… Genome language models (gLMs) have the potential to further understanding of regulatory genomics without requiring labeled data...

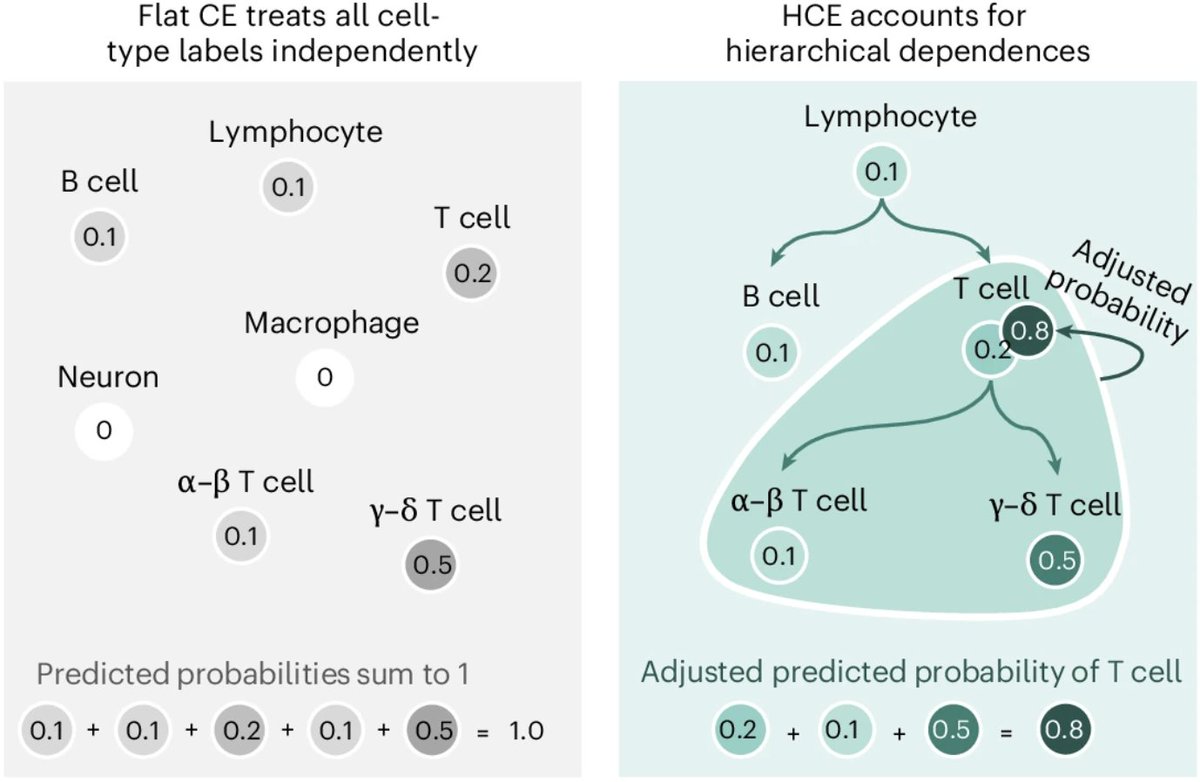
Excited to share our latest preprint, introducing the hierarchical cross-entropy (HCE) loss — a simple change that consistently improves performance in atlas-scale cell type annotation models. doi.org/10.1101/2025.0…




📢Out now! @sebacultrera, @davide_dascenzo, @avapamini, @peterswinter, @lorin_crawford, and colleagues present a hierarchical cross-entropy loss that improves performance of single-cell annotation models. nature.com/articles/s4358…

I’m deeply sad to learn of the passing of Dr. Bill Foege. Bill was a towering figure in global health—a man who saved the lives of literally hundreds of millions of people. He was also a friend and mentor who gave me a deep grounding in the history of global health and inspired me with his conviction that the world could do more to alleviate suffering. Bill and I often talked together about the future of global health. The legacy of his career is that many of the remarkable developments to come will have his imprint all over them. gatesnot.es/4sZmDqd

