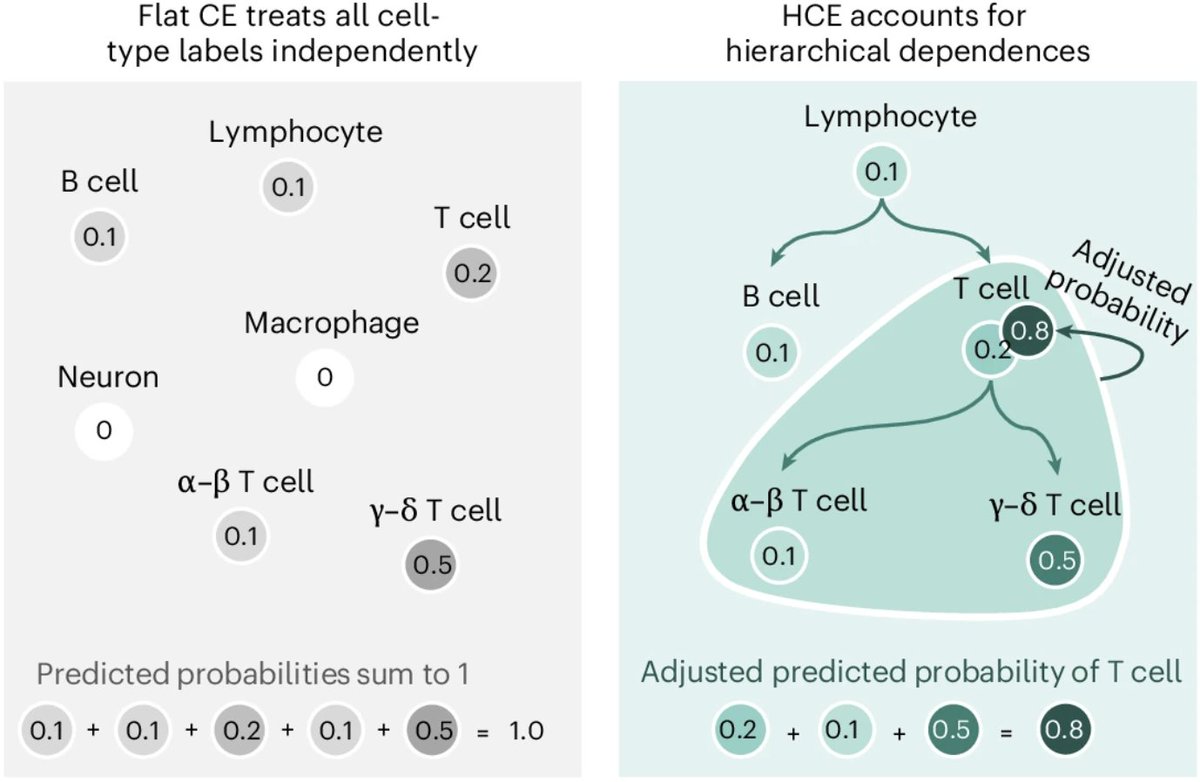
Excited to share our latest preprint, introducing the hierarchical cross-entropy (HCE) loss — a simple change that consistently improves performance in atlas-scale cell type annotation models. doi.org/10.1101/2025.0…
Sebastiano Cultrera di Montesano
45 posts

@sebacultrera
Postdoc at @Schmidt_Center within @Broadinstitute

Excited to share our latest preprint, introducing the hierarchical cross-entropy (HCE) loss — a simple change that consistently improves performance in atlas-scale cell type annotation models. doi.org/10.1101/2025.0…








𝘀𝗰𝗗𝗮𝘁𝗮𝘀𝗲𝘁 𝘃𝟬.𝟯.𝟬 𝗶𝘀 𝗵𝗲𝗿𝗲! 🚀 🎯 New features: • Native DDP support (no DistributedSampler needed!) • Built-in transforms (AnnData/HF/BioNemo) • Auto-config module 📝 Updated paper with theoretical guarantees! Details below 👇



Excited to share our latest preprint, introducing the hierarchical cross-entropy (HCE) loss — a simple change that consistently improves performance in atlas-scale cell type annotation models. doi.org/10.1101/2025.0…






📢Out now! @sebacultrera, @davide_dascenzo, @avapamini, @peterswinter, @lorin_crawford, and colleagues present a hierarchical cross-entropy loss that improves performance of single-cell annotation models. nature.com/articles/s4358…