Weize Xu
70 posts


Weize Xu
@Nanguage
Postdoc Scholar @Stanford BASE, Genetics | Build AI agent system for life science/biomedicine(PantheonOS: https://t.co/pdpUJ0jfMB)


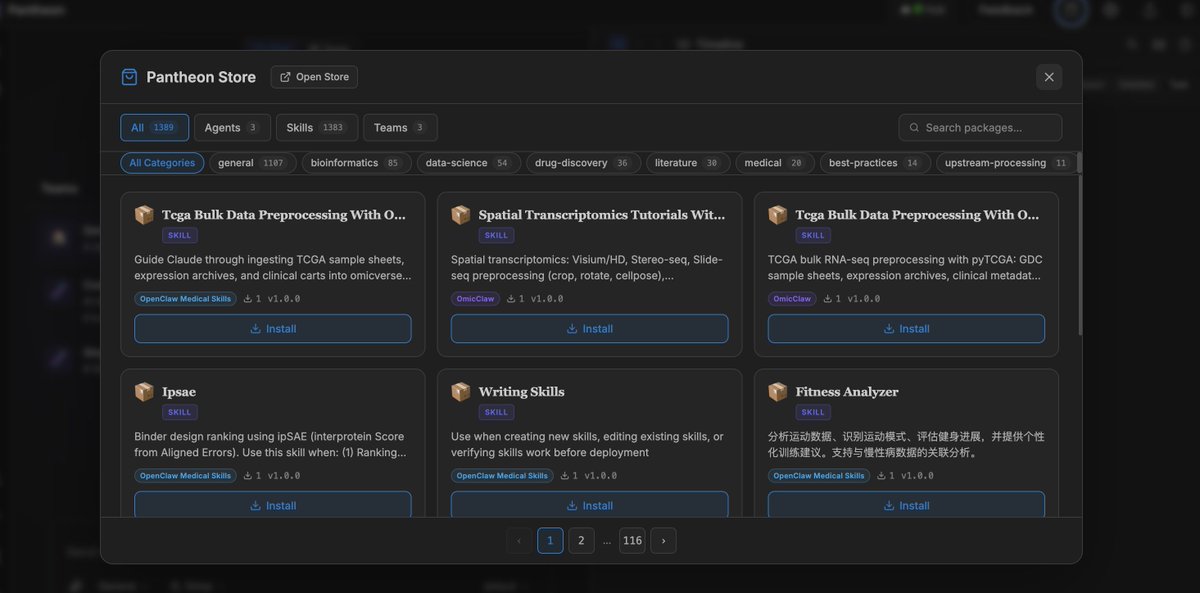
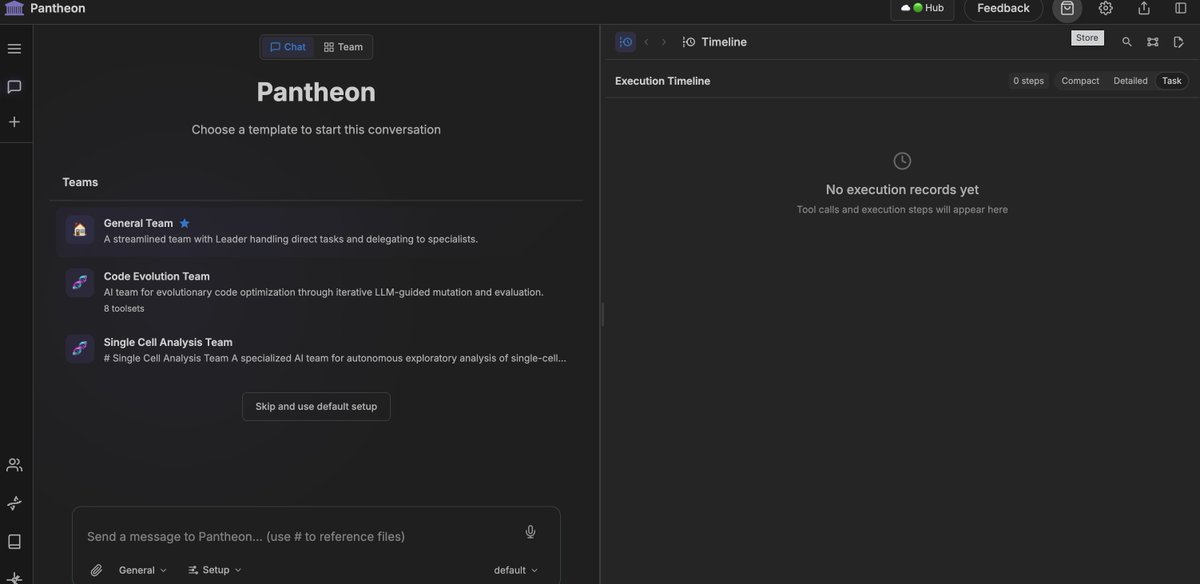


Pantheon AI Agent Store is now live with 1300 biomedical Skills and more! Pantheon Store is a new marketplace for biomedical AI Agents, Teams, and Skills. We launch the Store with 1300+ curated bio/medical AI capabilities, built by the incredible builders behind Claude Scientific Skills, ClawBio, OpenClaw Medical Skills, LabClaw, and PantheonOS ecosystem. You can install instantly in PantheonOS (UI or CLI) and build powerful scientific workflows right now. What you can do with Pantheon Store? It can: • Discover Agents, Teams, and Skills for genomics (especially single-cell and spatial genomics), pharmacology, medicine, and bioinformatics • Install Skills seamlessly into your existing workflows • Share your own Agents, Teams, and Skills with the community Thus, Pantheon-Store turns PantheonOS into a living ecosystem for scientific AI tools. We welcome you join the community! Upload your own components. Build together. Accelerate biomedical discovery!



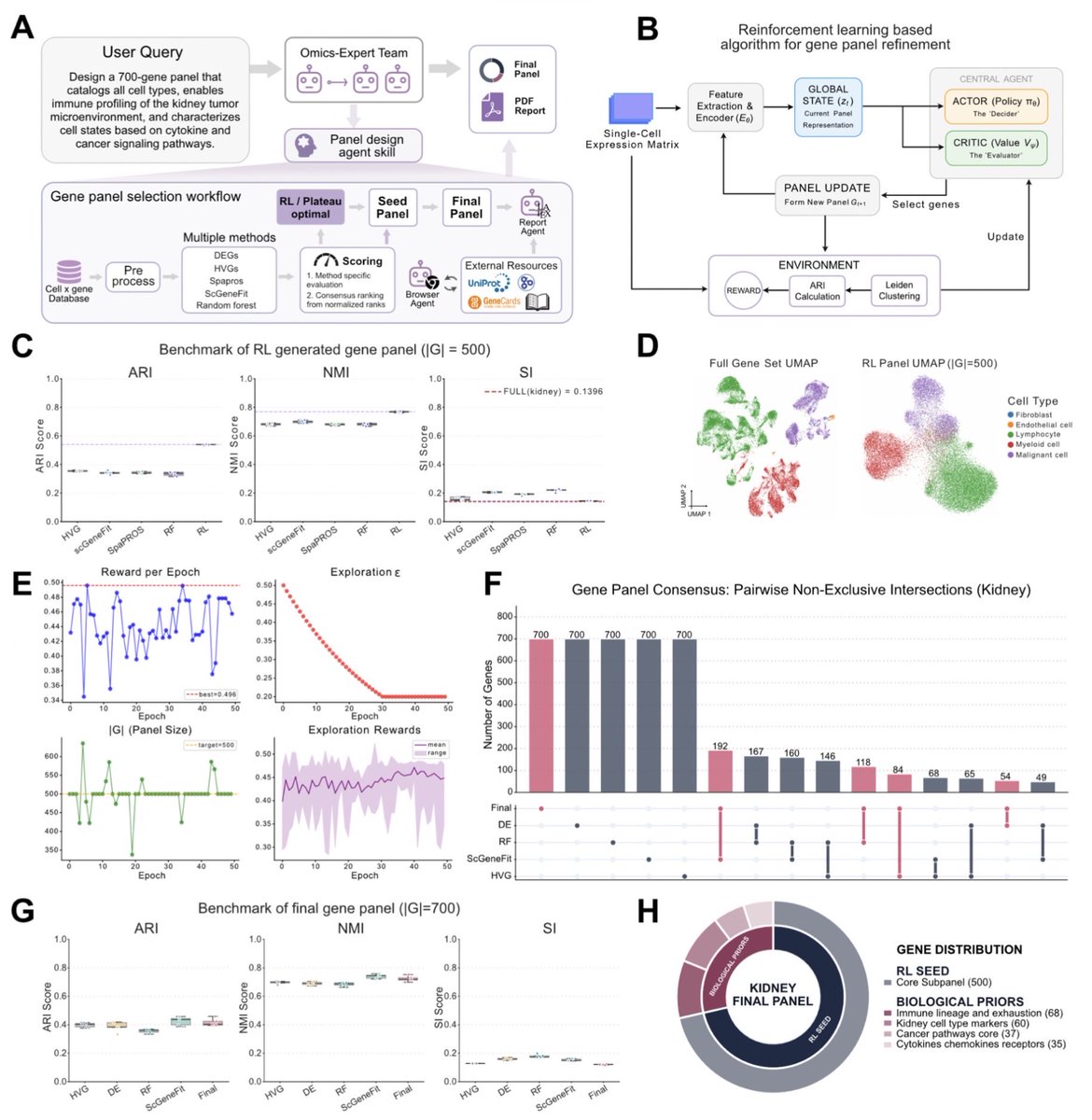










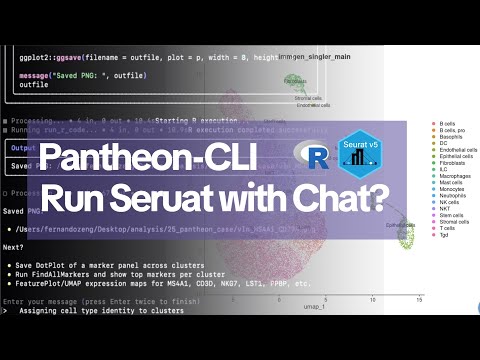
🚀 Introducing PantheonOS (pantheonOS.stanford.edu): A Fully Open-Source Agent OS for Science PantheonOS began as a research project in my Stanford lab and has since evolved into a vision to redefine data science in the era of AI—starting with computational biology, especially single-cell and spatial genomics. PantheonOS is a general agent platform built from the ground up. It is arguably the first distributed agent framework designed for scientific data analysis. 🔑 Key Features 1. Multi-Agent Collaboration – Built-in paradigms for distributed, cross-machine cooperation among agents and toolsets. 2. Native Toolset Support – Python, R, Julia, LaTeX, and more—designed for real scientific workflows. 3. Modular & Extensible – Developer-friendly design with shallow wrappers, plus LLM-driven toolset generation. 4. Evolvable Agents – Capable of evolving large-scale code projects to achieve superhuman performance (e.g., evolving upon the original Harmony [I Korsunsky, 2019, Nature Biotechnology] and Scanorama [BL Hie, 2019, Nature Biotechnology] implementations), and even evolving the system itself to adapt to new fields. 🎉 Stepwise Release Strategy We’re releasing PantheonOS in stages: Pantheon-CLI (today!), followed by Pantheon-Lab, Pantheon-Notebook, Pantheon-Slack, and more. 🌟 Pantheon-CLI Highlights - We're not just building another CLI tool. We're defining how scientists will interact with data in the AI era. - Open, Powerful, Python-First – The first fully open-source, endlessly extendable scientific “vibe analysis” framework. - Mixed Programming Magic – Combine Python, natural language, R, or Julia—seamlessly in the same environment. - PhD-Level Assistant – A command-line agent for complex real-world genomics and beyond, handling workflows at the PhD level. - Privacy by Design – Run entirely offline with local LLMs—your data never leaves your computer. ✅ Proven Applications (10 Demonstrations) Computational biology: 1. ATAC-seq: From raw reads to peak matrix 2. RNA-seq: From raw reads to expression matrix 3. Complex single-cell workflows (PhD-level) 4. Hybrid natural language + R for Seurat annotation 5. Learning from web tutorials + invoking single-cell foundation models 6. Cell segmentation on 10x Genomics HD Visium data And beyond: 7. Mixed Python & R programming examples 8. Molecular docking & structural analysis 9. Exploratory factor analysis for behavioral survey data 10. Customer segmentation & finance analytics 🌐 Learn More & Get Started Website: pantheonOS.stanford.edu Pantheon-CLI Documentation: pantheon-cli-docs.netlify.app GitHub Repo: github.com/aristoteleo/pa… 💬 Join our community: PantheonOS Slack: pantheonos.slack.com/ssb/redirect PantheonOS Discord: discord.com/invite/74yzAGYW