Angehefteter Tweet
Ufaq Bhat 🧕
1.1K posts


Ufaq Bhat 🧕
@UfaqBhat
Graduate student @IISERPune #Structural_Biology Don't be sad, Allah is with us
Srinagar Kashmir Beigetreten Ağustos 2015
447 Folgt401 Follower
Ufaq Bhat 🧕 retweetet
Ufaq Bhat 🧕 retweetet

Dr Pucadyil appointed as director of BRIC-NCCS, Pune indianexpress.com/article/cities…
@TPucadyil is currently Professor at @IISERPune
@DBT_NCCS_Pune
English
Ufaq Bhat 🧕 retweetet
Ufaq Bhat 🧕 retweetet

New Online! Overcoming the challenge of preferred particle orientation in cryo-EM dlvr.it/TRjcFR
English
Ufaq Bhat 🧕 retweetet

Piracy ke khilaf jung tab ladi jati h jab desh ameer ho. Bhai kisi ghareeb ka baccha 1-2 kitaab padh lega toh kya dikkat h. Importing first world priorities to third world countries.
Penguin India@PenguinIndia
Over 20,000 pirated books seized in Delhi through a collaborative crackdown led by Delhi Police with Penguin Random House India, @SimonSchusterIN & @HayHouseIndia - one of the capital’s biggest anti-piracy actions. Read the full report: penguin.co.in/newsroom/major…
Eesti

@Krishanpal_K @CellReports @MamatharaniV @mahipal_ganji @SamratLabMohali @IISERPune Congratulations to all authors. Fantastic work
English

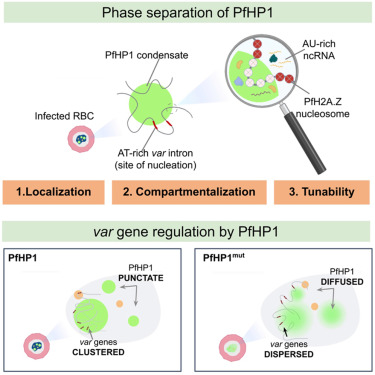
Excited to share our latest work @CellReports!
We show how PfHP1 phase separation is key to malaria virulence gene control.
cell.com/cell-reports/f…
Brilliant work by @MamatharaniV & the team. Thx @mahipal_ganji, @SamratLabMohali & Till Voss for a great collab! Thx @IISERPune.
English
Ufaq Bhat 🧕 retweetet

Built a tool for myself to easily trim/view/search/download/renumber protein structures for binder design tasks (even on the phone!)
wanted to share in case its helpful to anyone else
seq.design
English
Ufaq Bhat 🧕 retweetet

Some microbes carry a protein, called SNIPE, that "chops up" phage DNA as it's being injected into the cell.
This is a new mechanism for phage defense! CRISPR–Cas and restriction enzymes also evolved to fight against phages, but they work by recognizing sequences. SNIPE works, instead, by sensing "touch."
SNIPE is a protein with about 500 amino acids. After it's made by the ribosome, it latches onto ManYZ, two proteins which sit on the cell's inner membrane. (ManYZ is an importer; it brings mannose and other sugars into the cell.)
Once attached to ManYZ, SNIPE sits and waits for an invading phage. Some phages, including lambda, actually infect cells by pushing their DNA through this ManYZ channel. Lambda uses its "tail" to reach inside the protein channel, basically, and inject its DNA.
When this physical touch happens, though, SNIPE is waiting. As soon as the phage DNA starts entering the cell, and passes through ManYZ and SNIPE, it gets immediately destroyed.
This means that SNIPE is the first phage defense system discovered, so far, that uses spatial positioning at the injection site to destroy invaders.
But there are caveats, of course. If you untether SNIPE from ManYZ, such that it can freely diffuse through the cell, it will chew up the bacterium's genome. It is not a highly discerning nuclease! Also, SNIPE is not found in most bacteria. A prior pangenome study, which sequenced lots of different microbes, found that roughly a third of well-studied bacterial lineages had at least one member with a SNIPE-like protein. (For this paper, they just ported one of those homologs into an E. coli laboratory strain.)
And finally, because SNIPE's mechanism is tightly tied to ManYZ, it cannot be used to defend against phages that enter the cell through different routes. T4 phages, for example, inject their DNA straight through the cell membrane and into the cytoplasm, without interacting with ManYZ.
This is a nice basic science paper. Applications TBD. (Just remember that scientists figured out that bacteria had a phage defense system, called CRISPR-Cas, many years before it was repurposed into a gene-editing tool.)
P.S. The video below shows how cells with the SNIPE gene (middle row) kill invading phages, and thus continue growing and dividing. Empty vector (top row) refers to bacteria carrying a plasmid with no SNIPE gene; this is a control group. And SNIPE E414A refers to cells which received a mutated SNIPE gene, where the glutamate at position 414 has been changed to an alanine, thus destroying the protein's nuclease activity. These cells also die when they get infected with a phage.

English
Ufaq Bhat 🧕 retweetet
Ufaq Bhat 🧕 retweetet

Now on the Cover @PNASNews, Delightful to see @ganguly_akansha Journey from stress to regeneration, from leaf to root featured on cover. Akansha, you made it, very well done, congratulations.
English

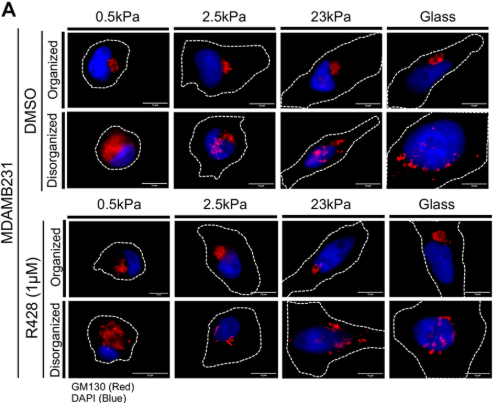
Arnav Saha @ArnavSaha19, Tushar Sherkhane @Tush96rkhane & Nagaraj Balasubramanian @AdhesionLab @IISERPune uncover a mechanosensitive pathway linking ECM stiffness to Golgi organisation & function in breast cancer cells.
journals.biologists.com/jcs/article/13…
Article: journals.biologists.com/jcs/article/13…
English
Ufaq Bhat 🧕 retweetet
Ufaq Bhat 🧕 retweetet
Ufaq Bhat 🧕 retweetet
Ufaq Bhat 🧕 retweetet

PIs & Group Leaders, do your trainees need to learn light microscopy and image analysis? Send them to us! Financial aid is available! Applications due Friday 1/30. meetings.cshl.edu/courses.aspx?c…
English
Ufaq Bhat 🧕 retweetet

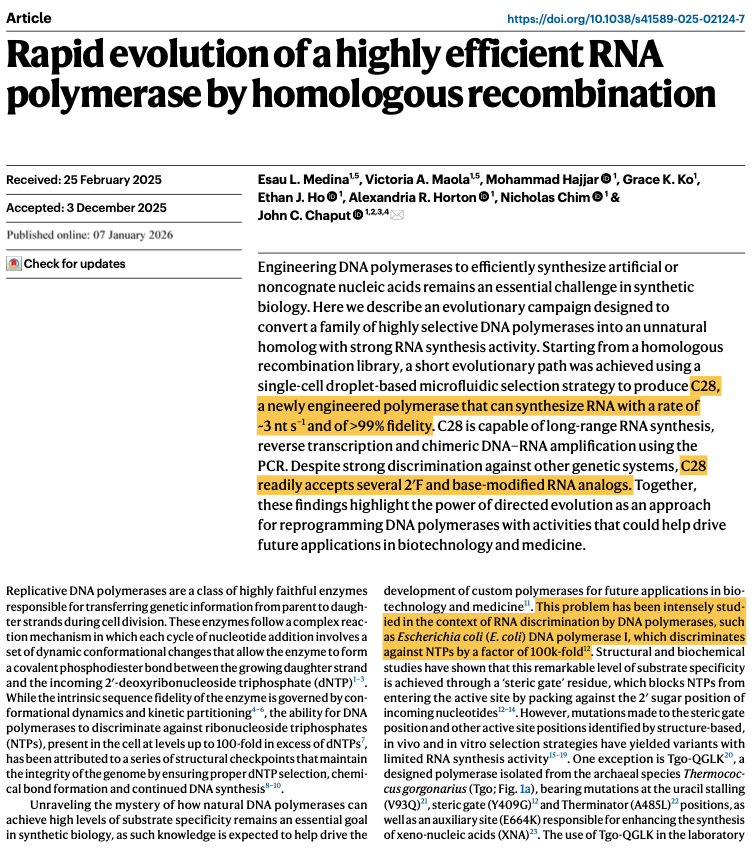
This paper is wild. After 3 rounds of directed evolution, they converted a DNA polymerase into an enzyme that can do:
- RNA synthesis
- Reverse transcription
- Synthesis of "unnatural" nucleotides
- Synthesis of DNA-RNA chimeras
One of the best papers I’ve read recently.
For context: In nature, it is DNA polymerase that takes a DNA sequence as a template and then copies it. These enzymes are crucial in replicating the genome for cell division, and they are EXTREMELY specific for DNA over RNA. This is key because RNA nucleotides are present in the cell at concentrations ~100x higher than DNA nucleotides, so the enzyme has evolved clever strategies to select one over the other.
RNA polymerases, for comparison, are the enzymes that take a DNA sequence as template and then convert it into RNA. They are involved in gene expression, for example.
To convert a DNA polymerase into an RNA polymerase (and all the other functions I mentioned earlier), the authors did a fairly straightforward directed evolution experiment.
First, they took four DNA polymerase enzymes belonging to various archaea. These DNA polymerases don’t check for DNA vs. RNA as stringently as other types of cells, so they’re a good starting point to evolve RNA polymerases. The authors inserted some targeted mutations into these enzymes, based on known mutations in the literature. For example, they swapped the amino acid at position 409 for a smaller amino acid, thus removing a “gate” that keeps RNA building blocks from entering the enzyme.
Next, they took the four genes encoding these DNA polymerases and cut them up into 12 segments each. They randomly stitched these 12 segments together — from the four different genes — to build millions of unique variants. Each shuffled gene was inserted into an E. coli cell.
Then, they grew up these cells (each carrying a unique polymerase) and put them into microfluidic droplets. A device isolates each droplet, lyses the cell open, and releases the polymerase. The droplet also contains RNA building blocks and a DNA template, encoding a fluorescent reporter. If the polymerase begins synthesizing RNA, it will produce a detectable signal. They screened about 100 million droplets in 10 hours of work, searching for those with a signal.
For each well that yields a fluorescent signal, the researchers isolated the DNA and sequenced it to figure out which polymerase it was. They repeated this 3x times, finally isolating a really excellent RNA polymerase variant which they called "C28."
C28 has 39 mutations compared to the wildtype enzymes. It incorporates about 3.3 nucleotides of RNA per second, with 99.8% fidelity. The crazy thing is that this enzyme can also copy DNA or RNA templates back into DNA (reverse transcription), or use chimeric DNA-RNA molecules as a template and amplify them. It is just a super versatile polymerase that can act on DNA, RNA, or modified nucleotides, to build just about anything.

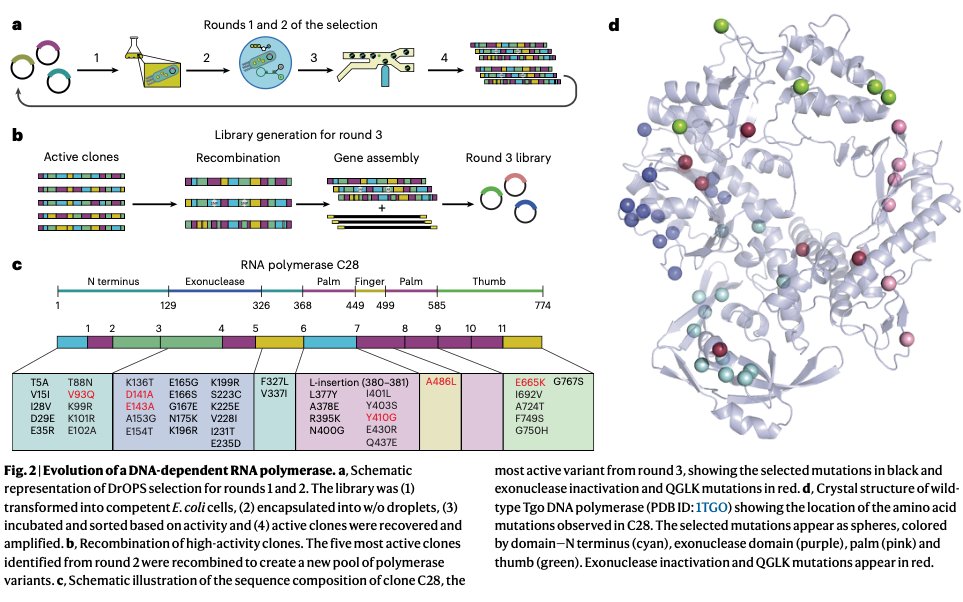
English
Ufaq Bhat 🧕 retweetet

"I sat in my supervisor’s office, red-faced and anxious, words tumbling out faster than I could control. For half an hour, I vented everything I had been holding in for months: the stress, the doubt, the sense that I didn’t belong. I was in the third year of my Ph.D., and a creeping fear had taken root that I wasn’t cut out for academia.
"I expected some kind of judgment or disappointment. Instead, my supervisor listened patiently, then calmly offered a line I’ll never forget: 'You are here to learn to ride a bicycle, not to invent a bicycle.' That one sentence landed softly, but it cracked something open."
Check out one of our top #ScienceWorkingLife essays of 2025: scim.ag/44IGRd7

English
Ufaq Bhat 🧕 retweetet
Ufaq Bhat 🧕 retweetet