Effi Kenigsberg retweetet

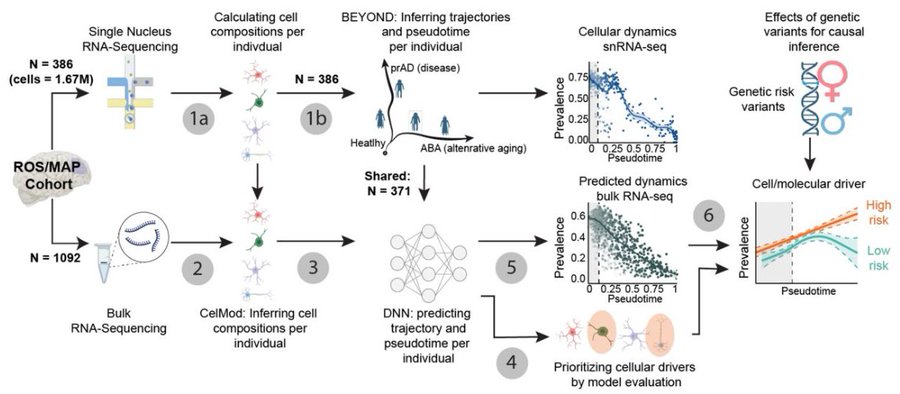
Presenting DeepDynamics, a joint effort by the @naomi_habib and @BarakRaveh Labs. We have built a deep learning framework that bridges bulk and single-cell RNA-seq to map the cellular cascades underlying Alzheimer’s Disease (AD) and aging. #alzheimer #deeplearning #ROSMAP
1/10
English