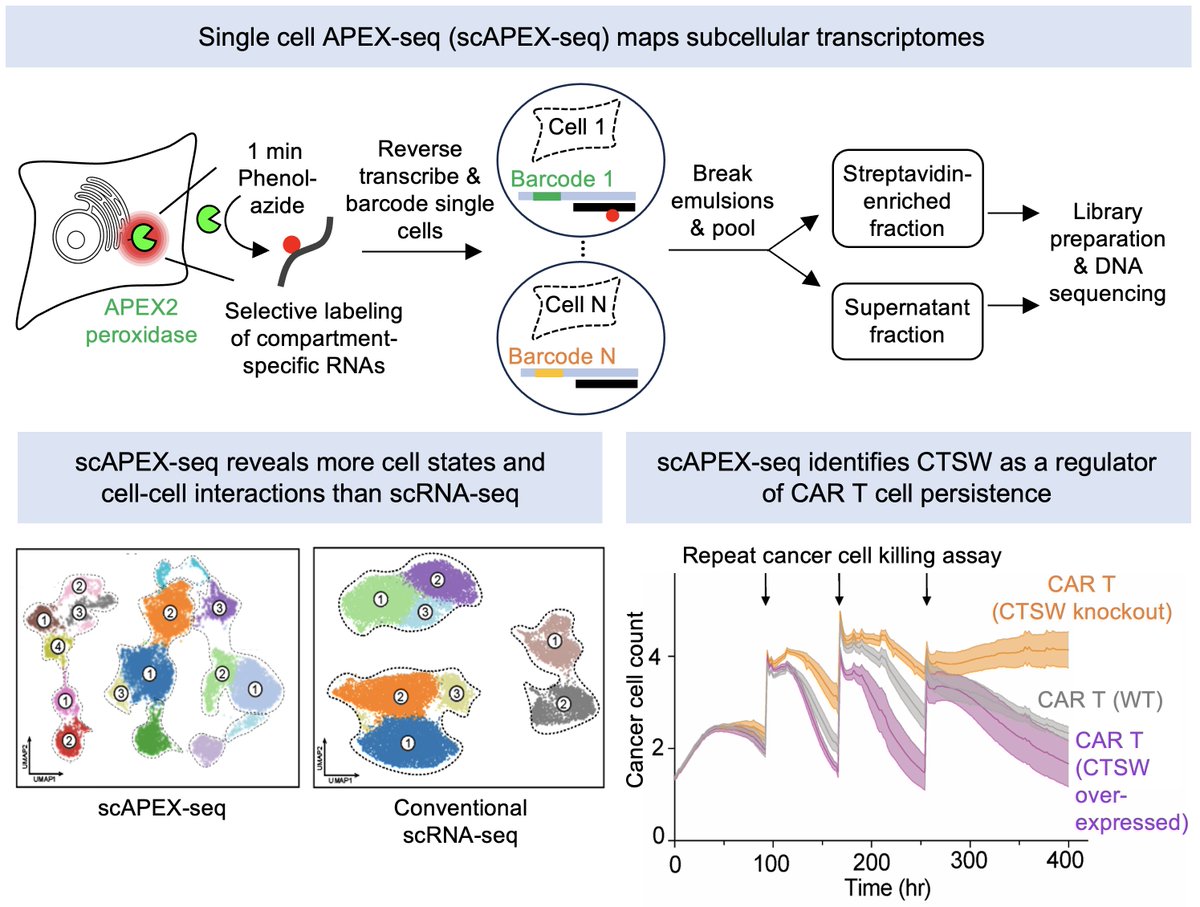
We are very excited to present the development of Zman-seq (“Zman”, Hebrew for “time”), the 1st technology that measures single-cell transcriptomes and physical time in vivo, led by @D_Birschenkaum, @CuriousKX, @FlorianIngelfi1, @AssafWeiner sciencedirect.com/science/articl…. (1/19)