Yuan Liu retuiteado
Yuan Liu
262 posts

Yuan Liu retuiteado

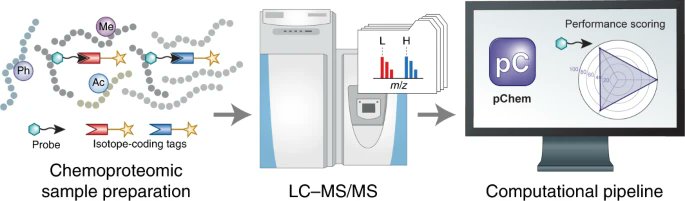
pChem, developed by @Yang_J_lab and coworkers, provides a computational pipeline for scoring chemoproteomic probes on their efficiency, modification homogeneity, and residue selectivity. Free to read link at rdcu.be/cR8SA
English
Yuan Liu retuiteado

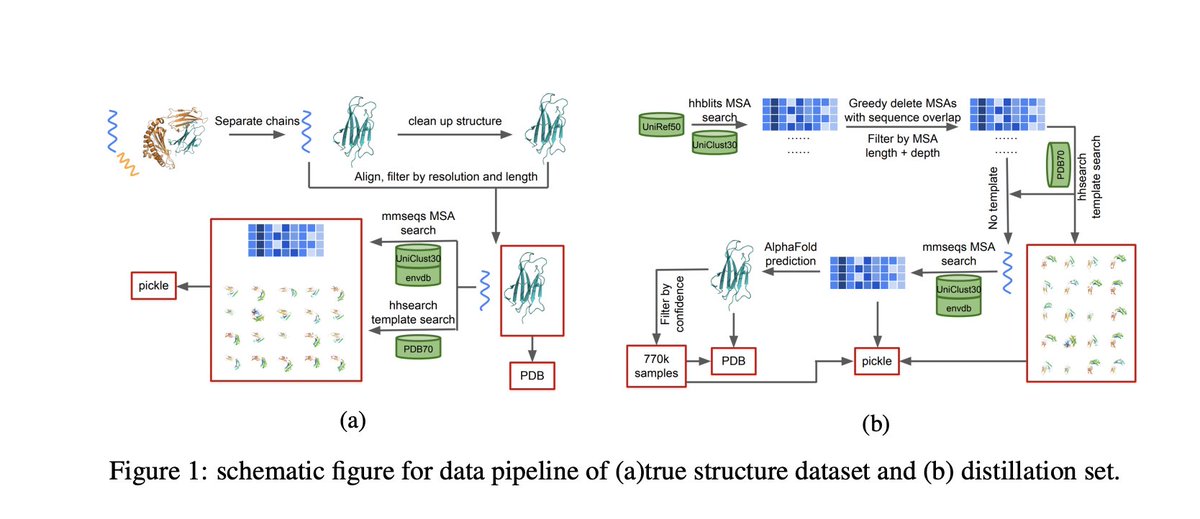
PSP: Million-level Protein Sequence Dataset for Protein Structure Prediction
abs: arxiv.org/abs/2206.12240
dataset consists of 570k true structure sequences (10TB) and 745k complementary distillation sequences (15TB)
English
Yuan Liu retuiteado

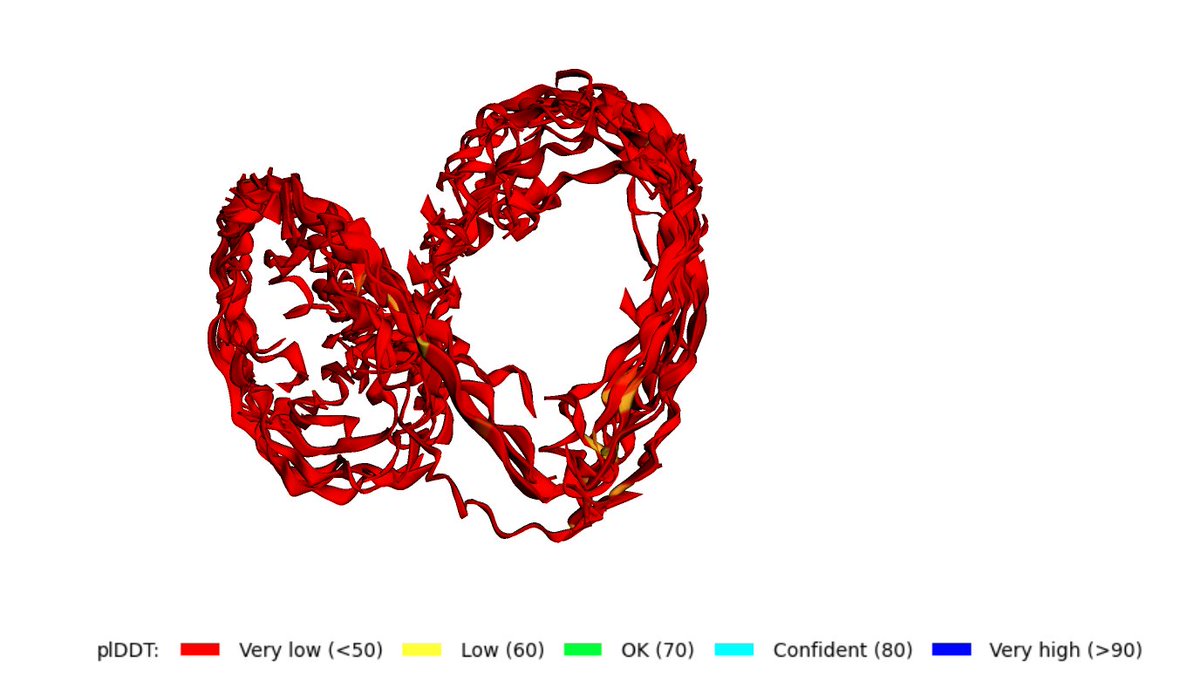
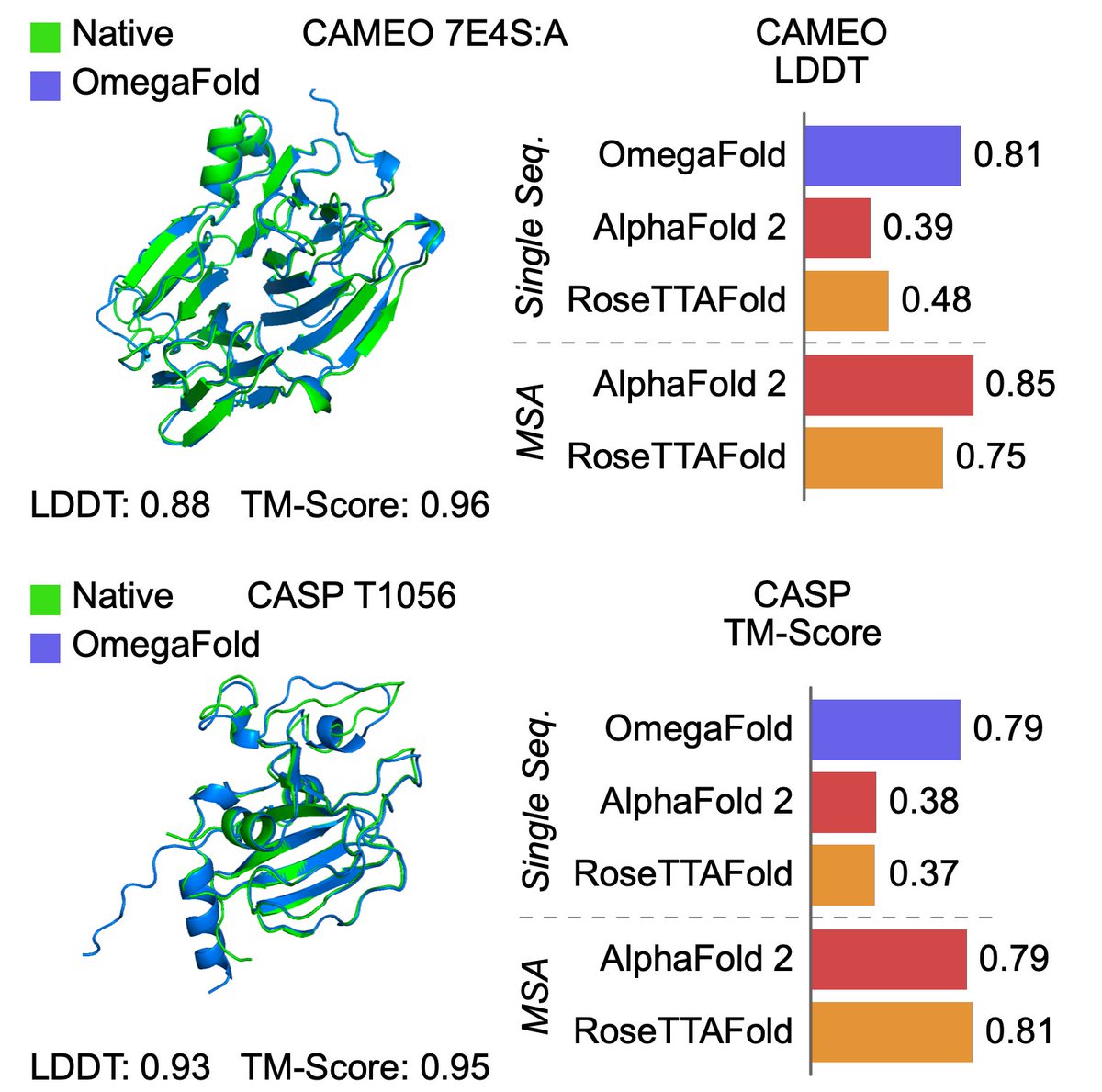
Protein structure can be predicted from a single sequence alone with high accuracy.
@HelixonBio team have developed OmegaFold, achieving performance similar to RF and AF2's MSA versions. Only a single sequence is given as input. 1/5
English

Supper fast!
Try MEQKPGTLMVYVVVGYNTDNTVDVVGGAQYAVSPYLFLDVGYGWNNSSLNFLEVGGGVSYKVSPDLEPYVKAGFEYNTDNTIKPTAGAGALYRVSPNLALMVEYGWNNSSLQKVAIGIAYKVKD with recycle = 12
Sergey Ovchinnikov@sokrypton
For my latest attempt at introducing proteins to students, I made a Google Colab Notebook that predicts proteins from a single sequence. I asked the students to tweak the sequence to get a helix or two helices or... (1/5) colab.research.google.com/github/sokrypt…
Svenska

Minor update to our decoy ranking manuscript. We (@jamesproney) add a plot showing that besides simply being able to rank decoys, AlphaFold can also be used to refine decoys:
biorxiv.org/content/10.110…
(1/5)

English
Yuan Liu retuiteado

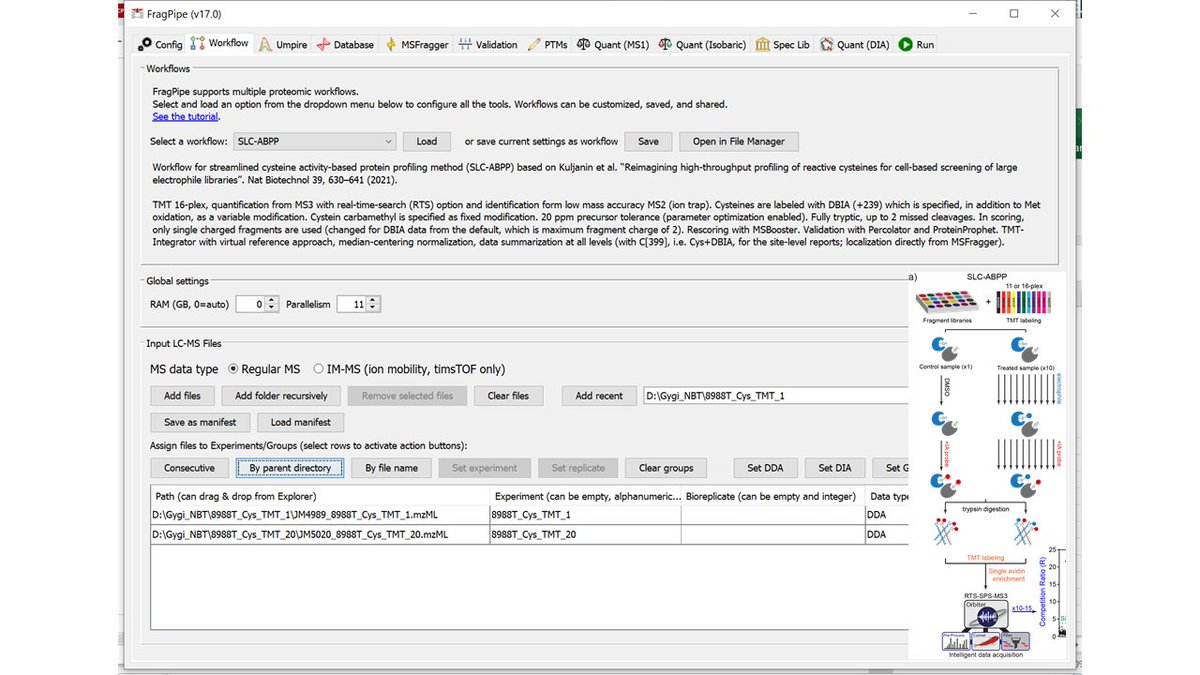
For those interested in activity-based protein profiling (ABPP), #FragPipe 17 comes with 3 workflows: classic isoTop-ABPP, SLC-ABPP (TMT16, DBIA tag nature.com/articles/s4158…), and isoDTB (chemrxiv.org/engage/chemrxi…). They can be easily customized. Let us know what else we should add.
English
Yuan Liu retuiteado

@UCDProteomics We are planning to create a #FragPipe "workflows repo" (including shared by others, e.g. from published papers). In addition to the more common ones we already include. So click 'Save workflow" and share with us, if you found settings that worked better than our default TMT-Ubiq.
English
Yuan Liu retuiteado

Our recent presentation on inverting structure prediction models for design is now public, for those that missed it! 😁
github.com/sokrypton/Cola…
Features @jueseph and new analysis from @JustasDauparas @KotaroTsuboyama @grocklin @minkbaek @sid_thesci_kid and others @UWproteindesign
Boston Protein Design and Modeling Club@ProteinBoston
For those that missed it, here is the recording! youtube.com/watch?v=2HmXwl… Special thanks to our speakers @sokrypton and @jueseph
English

Adding support for binder hallucination if anyone wants to try! (Code is very experimental, not intended for practical use... only use for art/science) 😀
colab.research.google.com/github/sokrypt…
English
Yuan Liu retuiteado

Just as convincing images of cats can be generated using artificial intelligence, new proteins can now be made using similar tools. In a new report in @Nature, we describe a neural network approach for “hallucinating” proteins with new, stable structures.
nature.com/articles/s4158…

English
Yuan Liu retuiteado

Huge thanks to our many users (fast growing userbase!) and the feedback we get via github.com/Nesvilab/FragP…. We just released #FragPipe 17.1 with minor fixes, including in MSBooster and in the DIA workflows, that affected a small number of users. If you are one of them, upgrade.
English
Yuan Liu retuiteado

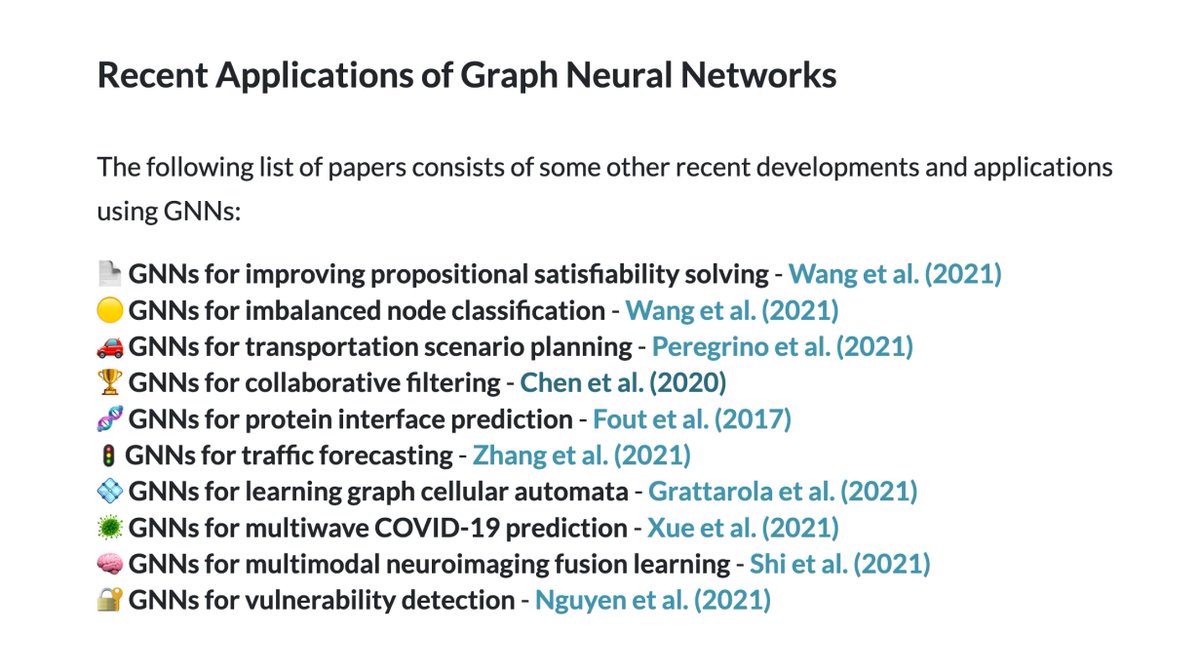
Graph neural networks are rapidly advancing progress in ML for complex graph data applications.
In this week’s newsletter, we show you recent uses of GNNs ranging from protein interface prediction to collaborative filtering! Read on below:
paperswithcode.com/newsletter/19/

English

@GoogleColab
Colab Pro+ seems to be broken. I upgraded last night hoping to increase runtime for a project... And the single Job was terminated after ~8 hours... now no more GPUs are available. 😭 (I had more resources w Pro... Now gotta wait till end of month to downgrade)
English
Yuan Liu retuiteado

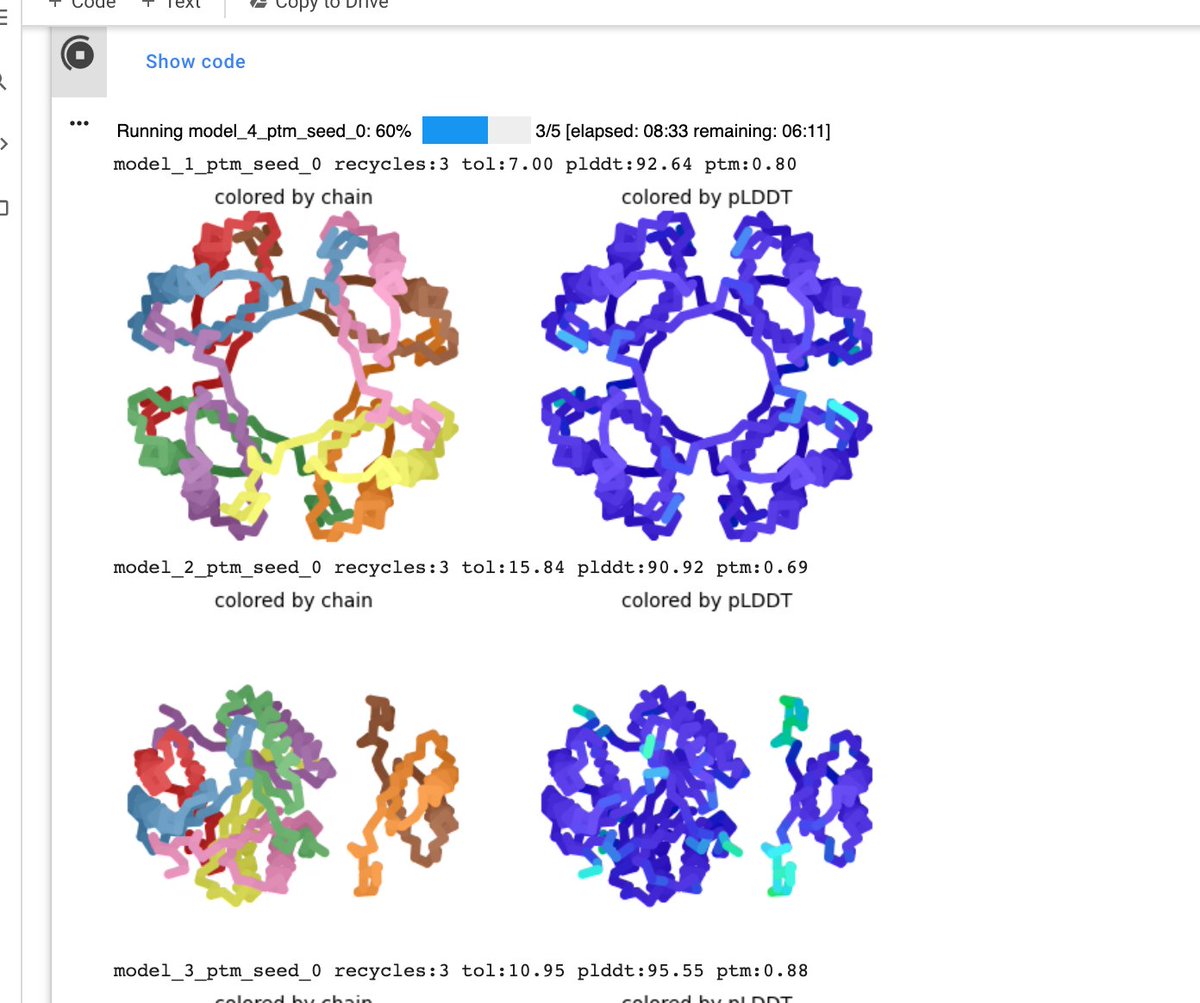
Impatient? Wanna see models as the notebook is running? 😎 See ColabFold AlphaFold2_advanced. Also, you can now use '/' to specify chain breaks. For example: A/C will modeled as A and C (could be used to trim disordered regions, or specify 2+ proteins) github.com/sokrypton/Cola…

English