

Sergey Ovchinnikov
3.8K posts


@sokrypton
Scientist, Assistant Professor @MITBiology, #FirstGen, ProteinBERTologist, 🇺🇦 No Human is illegal. Moving to: https://t.co/sow6IRD3jj




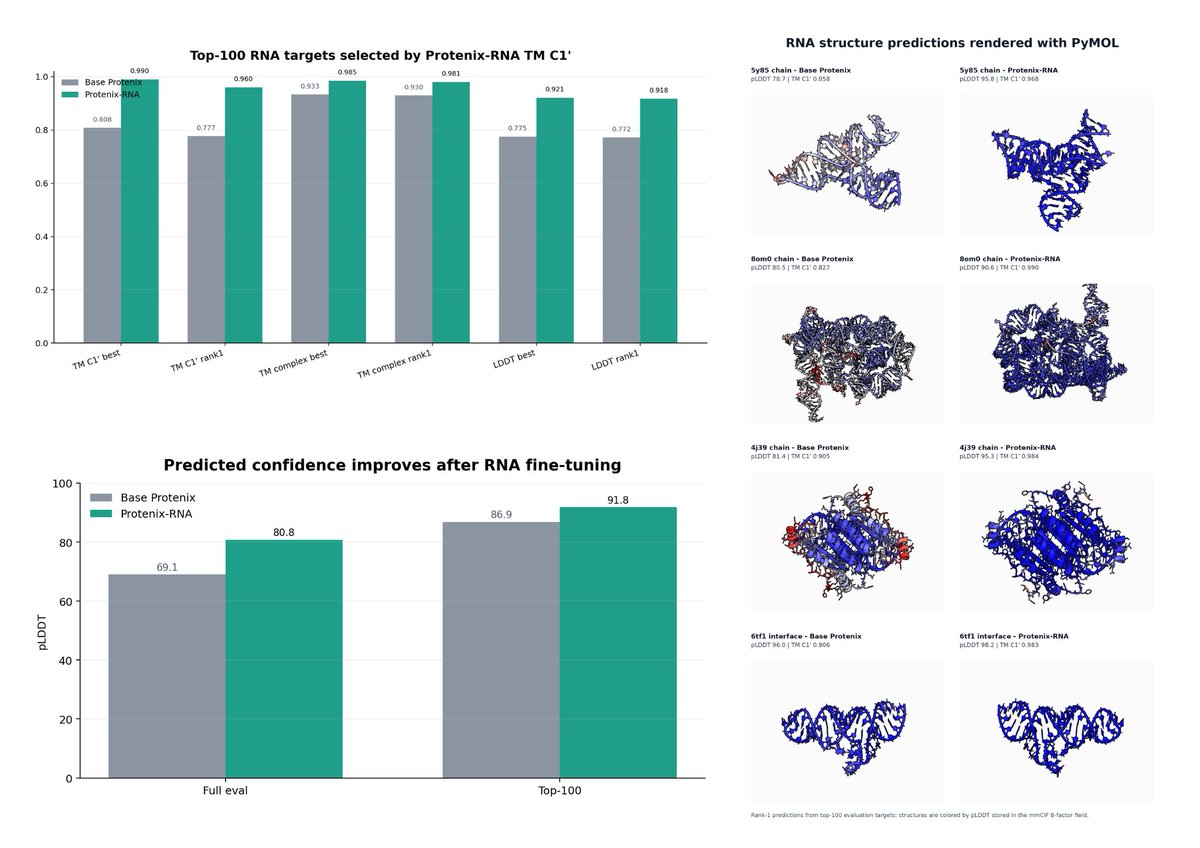
So our fine-tuning engine is giving us ~ 20% increase (full evaluation) in RNA folding task. Now starting second round of continual tuning.

We fine-tuned sequence- and structure-pretrained models on our large-scale stability data to predict absolute folding stability. The resulting SaProtΔG and ESM3ΔG achieved Spearman r = 0.88 and 0.87, with RMSE = 0.80 kcal mol⁻¹.






🚀 Excited to share our new work: Absolute Stability Predictor! 📊: forms.gle/4ZnXZSnTBvaykk… Built the MGnify Stability Dataset (1.8M+ measurements) and developed stability prediction models, together with @grocklin, @KotaroTsuboyama, @sokrypton, and teams.



We are releasing Carbon: a crazy fast DNA model Carbon is 275x faster than the next best model. So fast you can process the whole human genome on a single GPU in <2 days. Here are the tricks we used: When modelling DNA sequences a lot of the performance comes down to tokenizing the sequences in a smart way. BPE tokenizer struggle because there are no whitespaces and character (called base in DNA) level tokenizers waste a lot of compute on too many tokens. Carbon is built with a unique tokenizer: we split sequences in chunks of 6 bases, but during both training and inference we can work with single base resolution. That's similar to having word tokens but resolving them at the character level. All possible thanks to the DNA tokens unique structure. The architecture combined with the tokenizer makes the model 275x faster than the previous SoTA (Evo2) at this size. We built an interactive demo so you can explore how the model can generate DNA sequences, investigate the structure of genes, predict the effect of mutations, generate and fold proteins and even reconstruct parts of the tree of life. huggingface.co/spaces/Hugging…




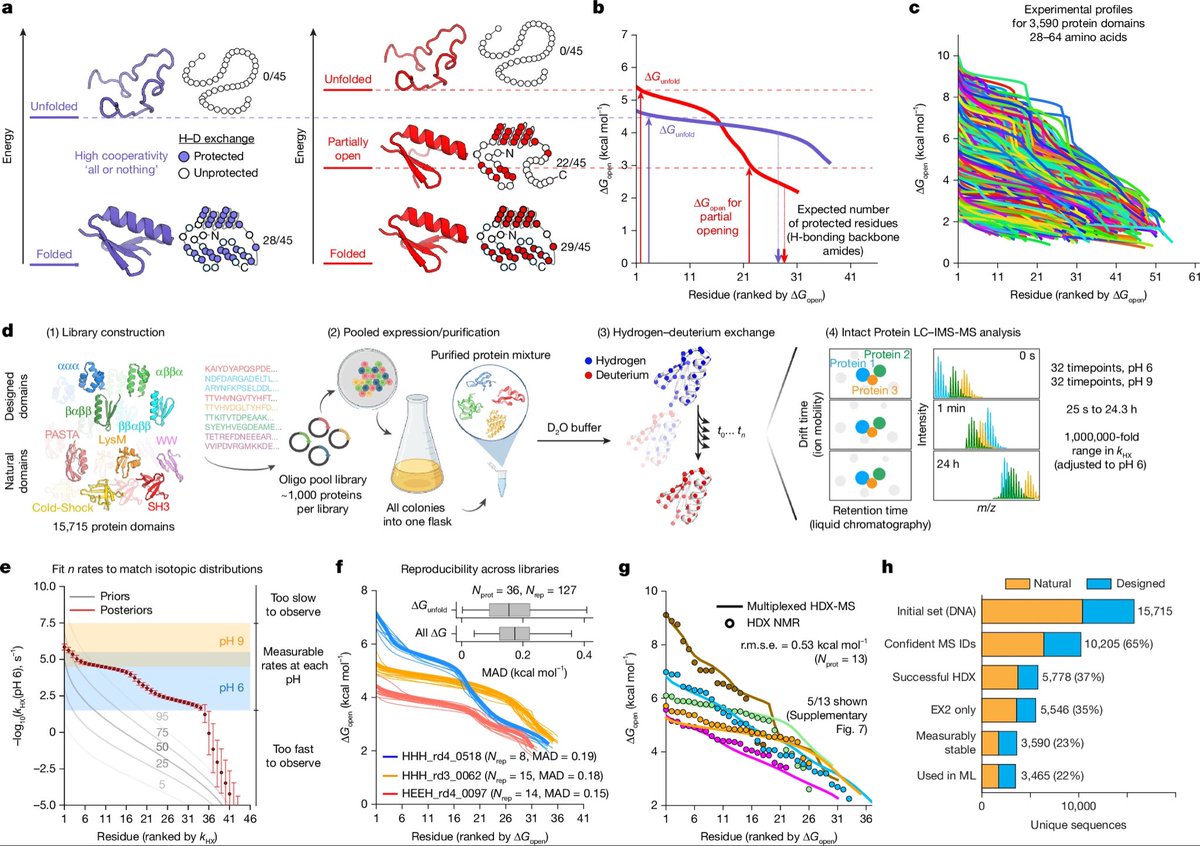
Thrilled to share our new paper where we introduce a multiplexed hydrogen–deuterium exchange MS (mHDX‑MS) method that can measure hundreds of protein domains’ conformational energy landscapes—all in a single experiment! biorxiv.org/content/10.110…

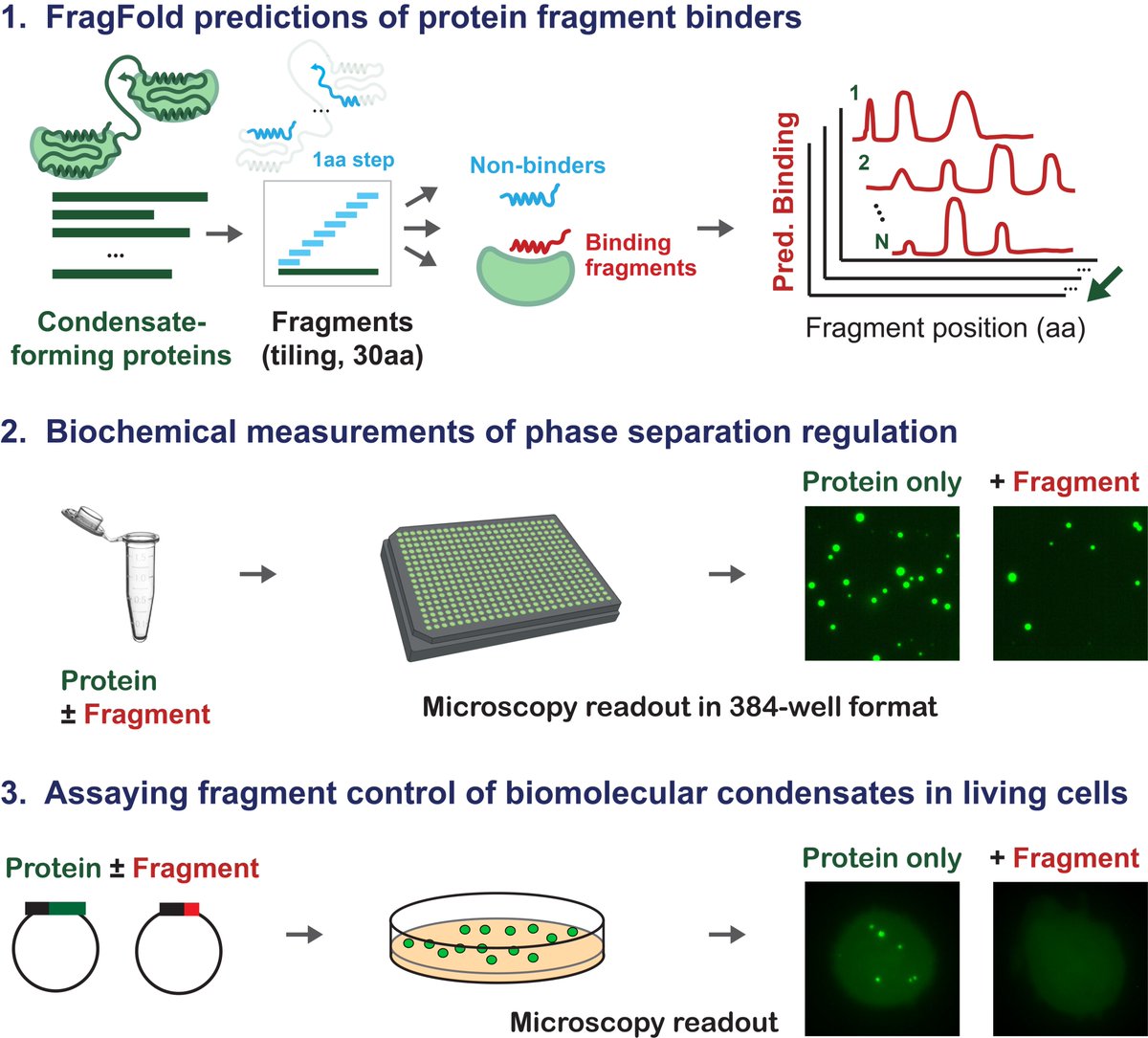






It’s estimated that the Protein Data Bank (PDB) cost around $13B to create. Alphafold was only possible because of it. If we want ML to solve biology, we should be funding the creation of databases and the development of new assay technologies. ML is nothing without data.