
Katie O'Mahony
1.6K posts


@forestsomewhere
PhD student @pharmabiotic @ucc | Microbes, Biofilms and Antimicrobial Resistance | All views my own | @[email protected]






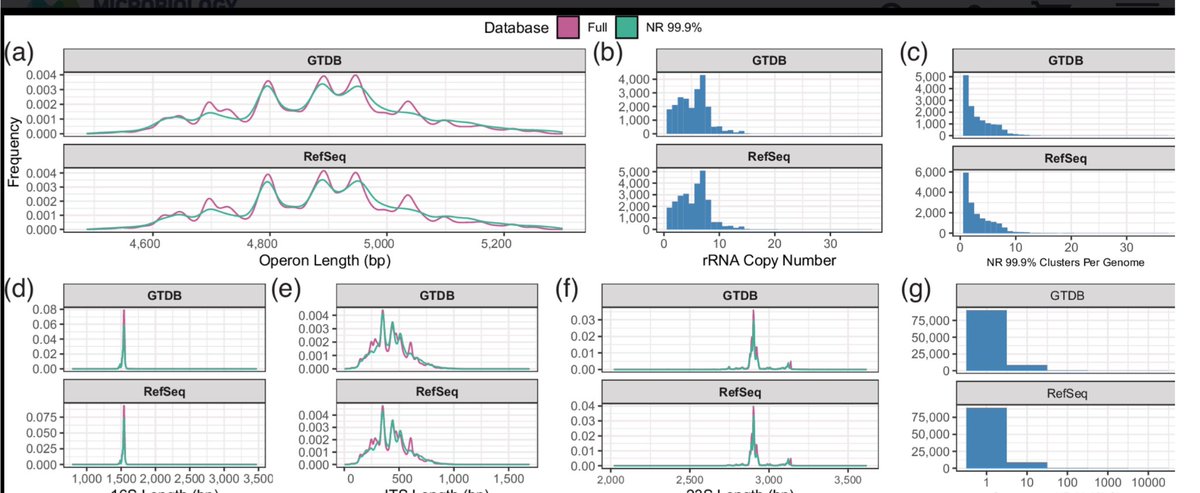

Some weekend reading…great to see GROND, our new database for long-read amplicon sequencing of the ribosomal operon, is out. Congrats @calumari_walsh @meghana_srini @VanSinderenLab @pauldcotter @tstinear microbiologyresearch.org/content/journa…

@rrwick, myself and the team (@JuddLmj @tstinear @linsalrob & Sarah) have recently looked at the effect of depth on short-read polishing almost-perfect Nanopore bacterial genomes biorxiv.org/content/10.110… (1/7)

















Researchers in Ireland are protesting today because our wage is set at the poverty line. Not even minimum wage. This needs to change in the 2023 budget. #PGRsDeserveBetter @paulmurphy_TD @RBoydBarrett @pb4p @hea_irl @DeptofFHed @SimonHarrisTD
