עודד | oded
2K posts


עודד | oded
@_odedf
Building the data layer for biology @strandaibio | YC W26






🚨: Japanese scientist Patrick Soon-Shiong has designed a treatment that activates body's natural killer cells that fight against cancer cells. Its approved in the U.S. and now Saudi Arabia has also approved it for its public.









Immunotherapy works. The problem is how we decide who gets it. Right now, a patient gets a single biopsy stain from one location in the tumor, and if it reads negative, they're told immunotherapy isn't an option. That test is a snapshot. Tumors are dynamic. They actively remodel the immune environment around them, suppress immune cell infiltration, and even excrete checkpoint proteins to shut down the response at distant sites. We see patients regularly who were told immunotherapy wouldn't work for them based on that one result. Making real spatial immune profiling accessible from a slide every cancer patient already has on file means fewer people being turned away from treatments that could help them. That's what matters here.




Our live tissue clearing paper is out in @naturemethods! We achieved optical clearing of mammalian brain tissues without compromising normal neuronal function. Big congrats to @Shigenori774 and our wonderful collaborators! 🎉 nature.com/articles/s4159… (1/10)

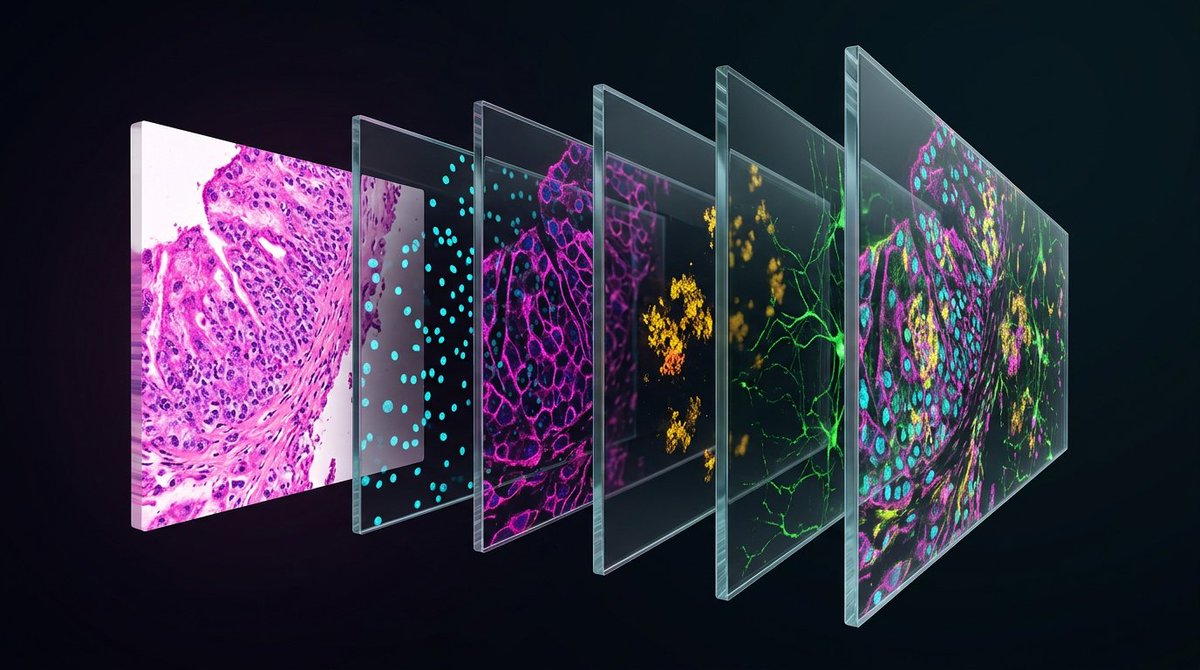
Introducing POSTMAN. POSTMAN computationally predicts spatially-resolved protein expression directly from routine H&E pathology slides. What used to require thousands of dollars and days of specialized lab work, you can now get from the slides already sitting in your archive.