Soham Mukhopadhyay
445 posts


Soham Mukhopadhyay
@SohamBio
Plant Pathogen Interaction | Researcher & Adjunct Faculty @UNM | Associate Editor @MPMIjournal🌐 https://t.co/m5PGg432yA | Try https://t.co/BI10lkHdvp



A clubroot pathogen PBS3-like effector manipulates hormonal crosstalk to alter root morphology during colonization biorxiv.org/content/10.648… #biorxiv_plants









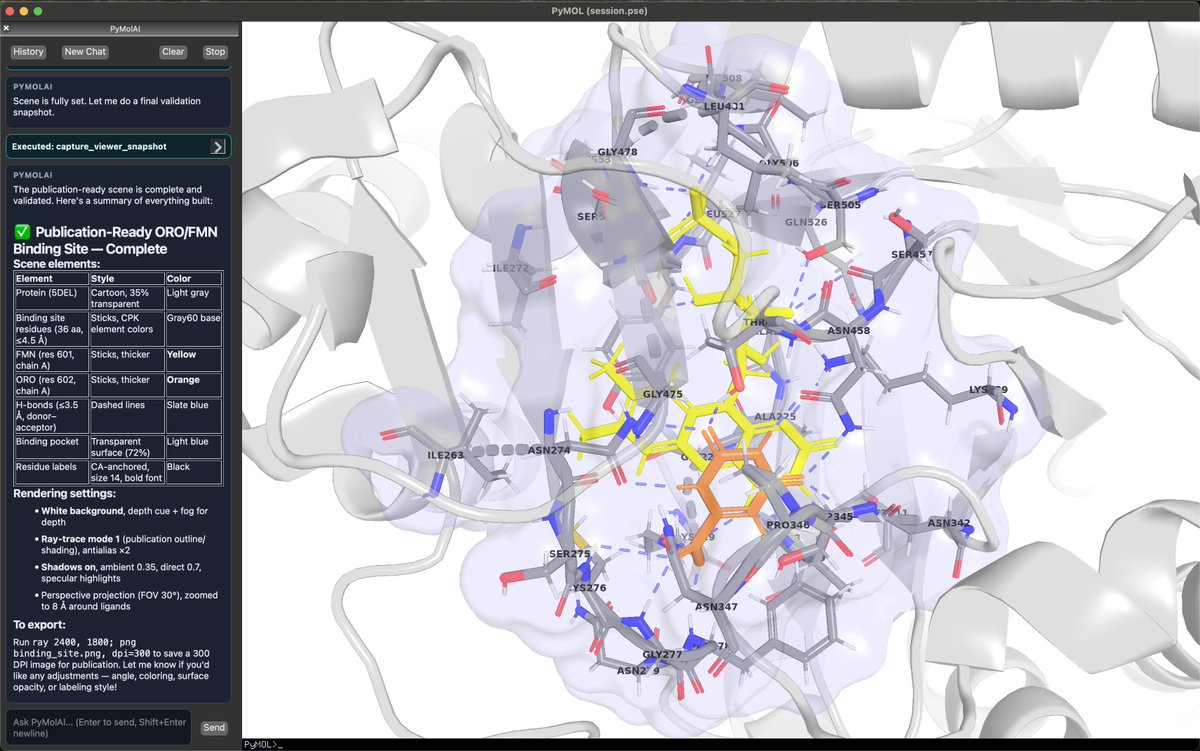
I am Open-Sourcing PyMolAI! Meet PyMolAI, an AI agent that can talk to your protein structures. Built on top of PyMOL, PyMolAI lets you interact with your structures in plain language. Whether you're: - Analyzing protein structures - Aligning complexes - Creating publication-ready figures - Or running design workflows PyMolAI interprets your request, executes the necessary PyMOL commands, and manages the workflow for you. It integrates with @OpenBioAI APIs, giving you access to tools like Boltz, ProteinMPNN, and BoltzGen — directly from your PyMOL session. It has local chat history with session syncing, so you can pick up exactly where you left off.

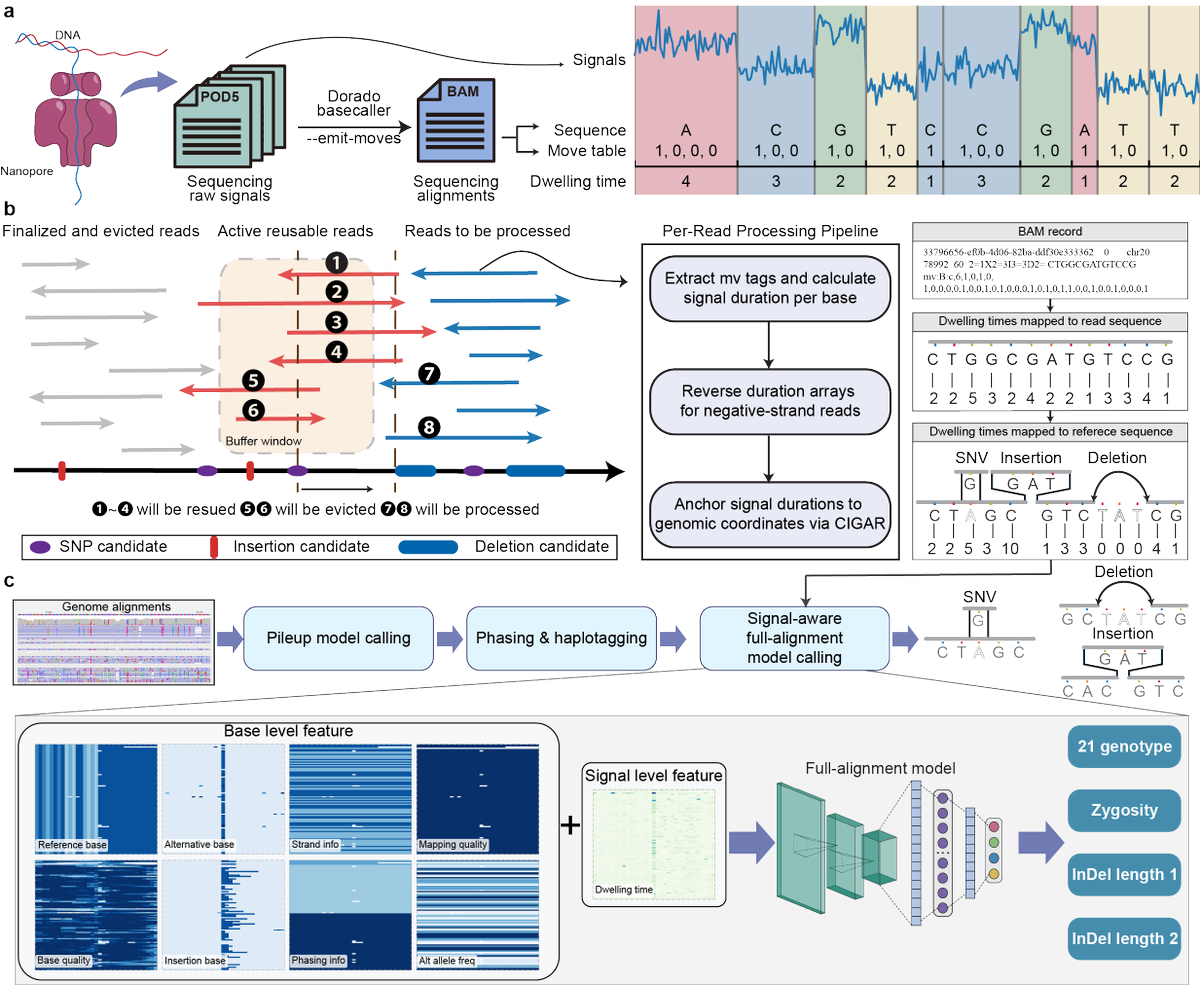


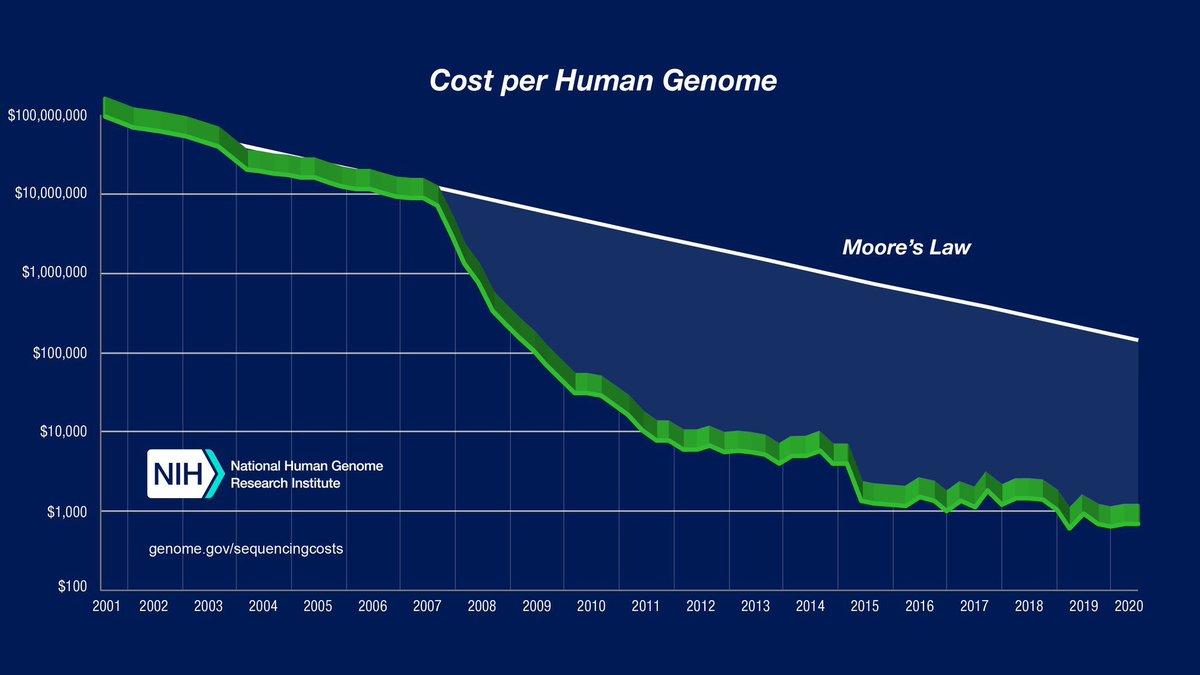
The $100 whole human genome sequence finally reached! @ElemBio sandiegouniontribune.com/2026/02/19/scr…


Thrilled to announce that I am joining the Editorial Board of Molecular Plant–Microbe Interactions (@MPMIjournal) as a Senior Editor! 🌱🦠 Grateful for the opportunity to serve this vibrant research community and help amplify impactful science 👏 🔗 apsjournals.apsnet.org/page/mpmi/edit…