固定されたツイート

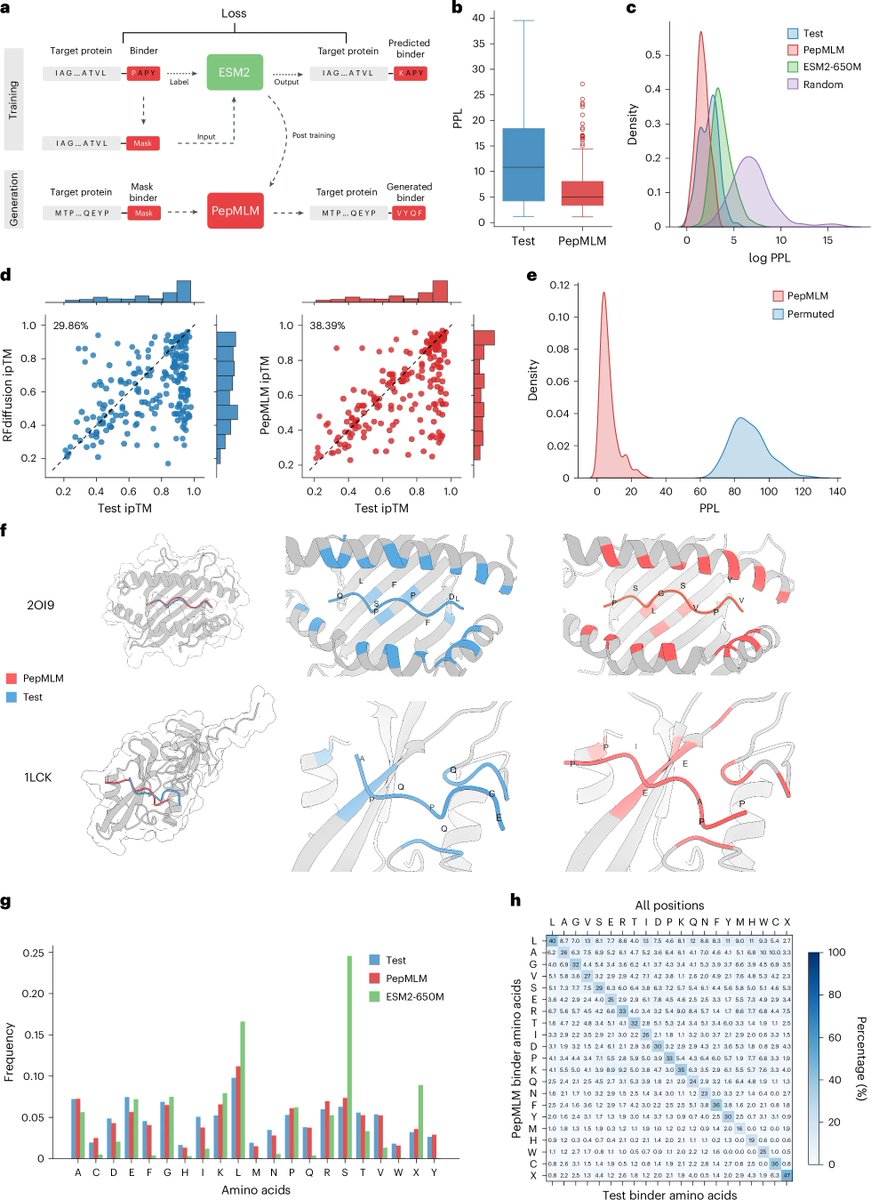
🚨 New Paper Alert
Sialic acid identity shapes host tropism in sialoglycan-binding viridans group streptococci
🔗 jbc.org/article/S0021-…
📍 @jbiolchem @VanderbiltU @ORNL
#Glycoprotein #Sialoglycan #CompChem #microbiologylove
English
Rupesh Agarwal
2.2K posts

@rupeshagarwal
रूपेश अग्रवाल| Computational Chemistry/Biophysics Scientist | Jack of all, Master of some and PhD of one😀











RFdiffusion2 is now live! github.com/RosettaCommons… You can now design proteins, and in particular enzymes from just partially defined amino acid side chains, and without defining their sequence position or order! Great job @r_krishna3 @woodyahern @dougtischer and everyone else!