고정된 트윗
Ben Busby!
6.8K posts

Ben Busby!
@DCGenomics
I work for NVIDIA and help out at CMU, JHU and Stanford in an effort to make Bioinformatics Better. Everything I post is my personal opinion. You're awesome!
Pittsburgh, PA 가입일 Temmuz 2013
1.4K 팔로잉2.1K 팔로워
Ben Busby! 리트윗함

Pharmaceutical giant Roche announced it is launching a new large-scale “AI factory” with thousands of the latest Nvidia chips to accelerate the development of new drugs and diagnostics.
Roche said the company has ramped up its AI capacity by purchasing 2,176 Nvidia Blackwell GPUs, which will be deployed across the U.S. and Europe.
That brings its total GPUs to more than 3,500, which it claims is the greatest number owned by any pharmaceutical company.
Roche, which made the announcement in conjunction with Nvidia’s GTC conference in San Jose, hopes to use that technological firepower to discover, develop, manufacture and commercialize therapies faster.
Learn more: forbes.com/sites/amyfeldm… (📸: Richard Morgenstein 2021)

English
Ben Busby! 리트윗함

Academic friends who work on AI x Science, consider applying to the Google.org Impact Challenge on AI for Science before Apr 17. The funding amount ($500k-3M) is substantial to help push research for a moonshot. I'm happy to help provide feedback on some proposals, ff to email me !
English
Ben Busby! 리트윗함

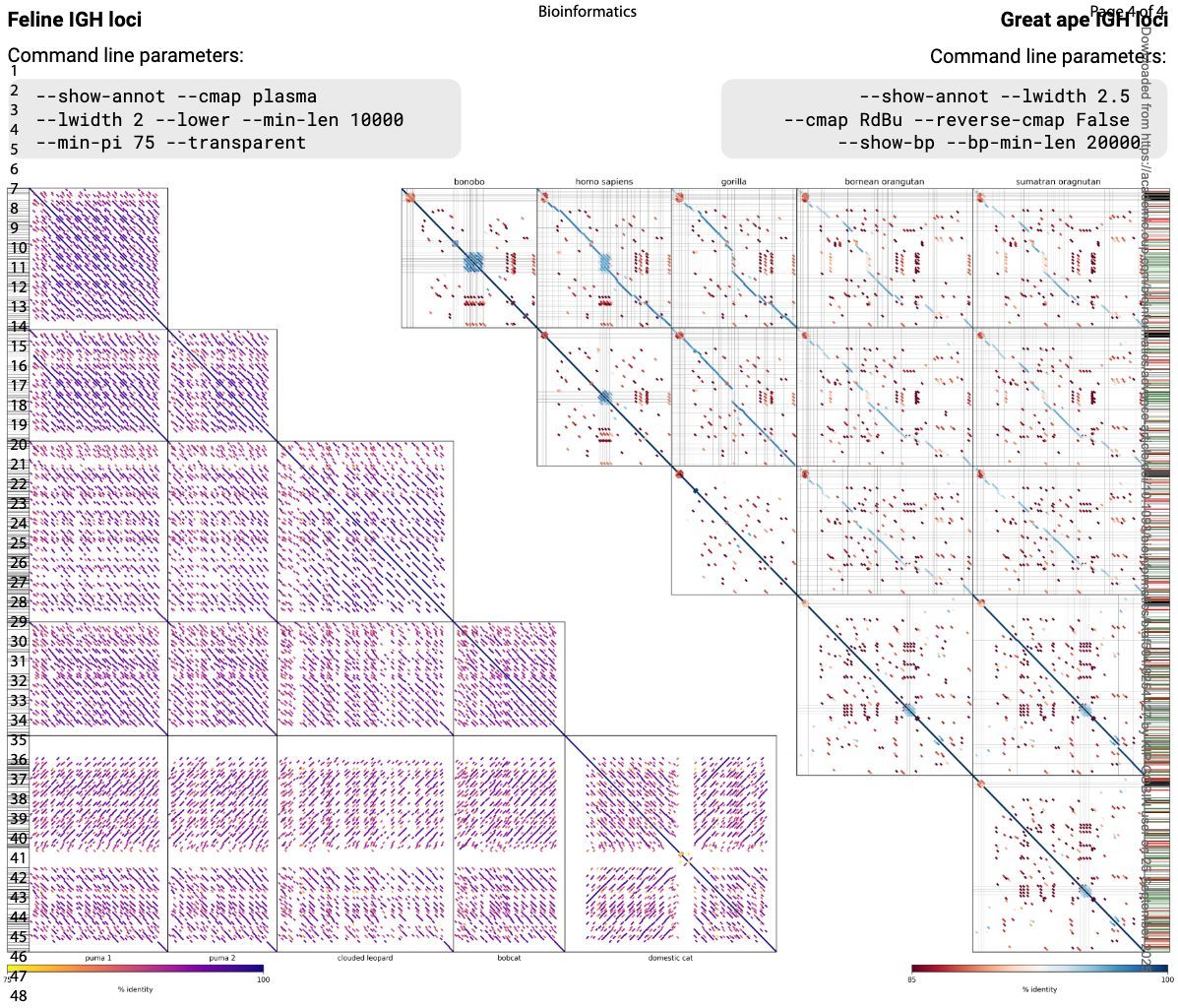
PatchWorkPlot: simultaneous visualization of local alignments across multiple sequences academic.oup.com/bioinformatics… 🧬🖥️🧪
English
Ben Busby! 리트윗함

AI to understand the language of life.
Our review @NatureBiotech just published
nature.com/articles/s4158…
Free access rdcu.be/e89lQ
@VishRao5 @serena2z @BrianPlosky @pdhsu @BoWang87 @james_y_zou @marinkazitnik @pranavrajpurkar
English
Ben Busby! 리트윗함

Abstract deadline for CSH Symposium on AI in Biology is next week, Mar 26!
This frontier AI x Bio meeting will span many scales of biology, from proteins and genomes to cells, medicine, and agents, with 50+ invited speakers who are leaders in the field.
meetings.cshl.edu/abstracts.aspx…
English
Ben Busby! 리트윗함

Introducing PerturbAI.
Today we announced our emergence from stealth with the release of the world’s largest in vivo CRISPR data engine, interrogating the effects of thousands of genetic perturbations across 8 million cells throughout the whole brain.
This dataset represents a new category of biological data: organism-level, circuit-resolved causal genomics leading to novel targets and therapeutics.
By combining scalable in vivo CRISPR perturbation with AI, we model biological systems at unprecedented resolution and simulate therapeutic interventions before committing to expensive downstream development.
We’re grateful to our collaborators at @NVIDIAHealth and @10xGenomics for helping make this landmark dataset possible.
Read More: perturb.ai/news
#CRISPR #AI #DrugDiscovery #FunctionalGenomics #Biotech
English
Ben Busby! 리트윗함

Since it’s GTC week - good time to reintroduce what I do at @nvidia. I manage the investment portfolio of NVIDIA Ventures, working with startups across frontier model builders, techbio, robotics, AI infrastructure, materials discovery, energy, quantum and healthcare. I frequently draw on my training as a scientist and product manager for my job.
My primary goal is to drive tech and ecosystem integration between NVIDIA and portfolio companies. Which can eventually lead to successful long term relationships.
I am most interested in areas where AI translates into real world systems, especially in biology, robotics, and other scientific applications.
More on here - nventures.ai
English
Ben Busby! 리트윗함

Eduomics: a Nextflow pipeline to simulate -omics data for education academic.oup.com/nargab/article… 🧬💻🧪 github.com/lescai-teachin…



English
Ben Busby! 리트윗함

📢BioHackathon Europe 2026 Call for project proposals is now open!
Submit your #bioinformatitcs project for #BioHackEU26 to meet with other #bioinformaticians to drive the project forward.
📥 Submit your proposal by 15 April: biohackathon-europe.org

English
Ben Busby! 리트윗함

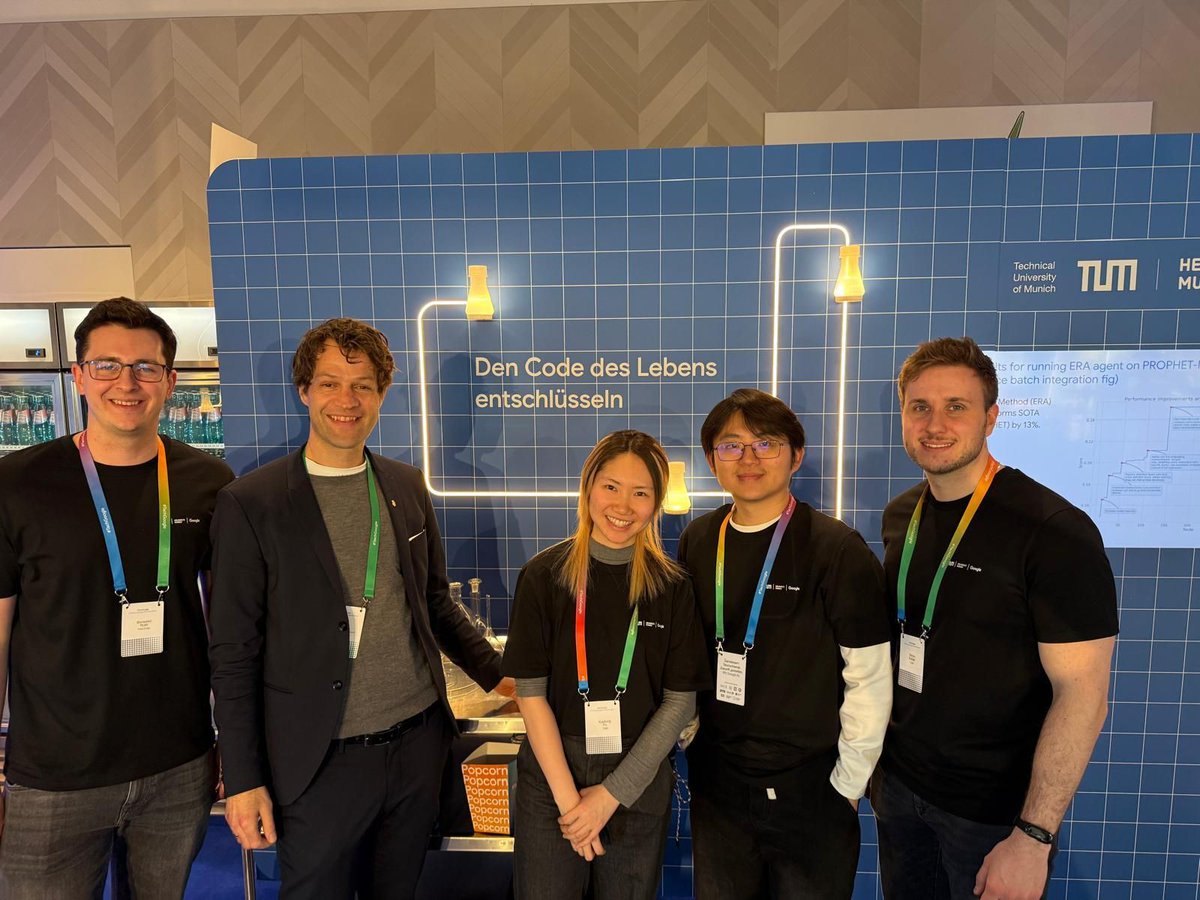
It's hard for me to contain my excitement for the work fo the Digital Biology Team @NVIDIAHealth #NVIDIAGTC this year. I can't WAIT for you all to see what we have been up to. I've got a detailed talk on Wednesday morning where I'll go through all the details on behalf of the team, but there will be lots to discover tomorrow and Tuesday too!
nvidia.com/gtc/session-ca…
English
Ben Busby! 리트윗함

You have my attention... 👀
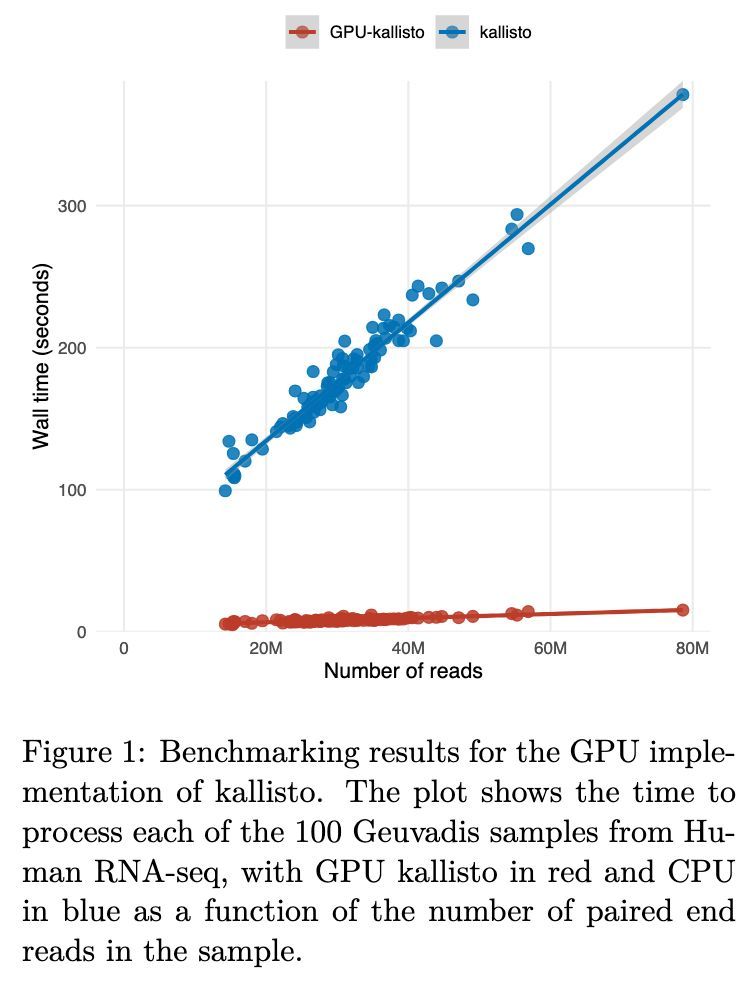
RNA-seq analysis in seconds using GPUs biorxiv.org/content/10.648… 🧬💻🧪 github.com/pachterlab/kal…
English
Ben Busby! 리트윗함
Ben Busby! 리트윗함

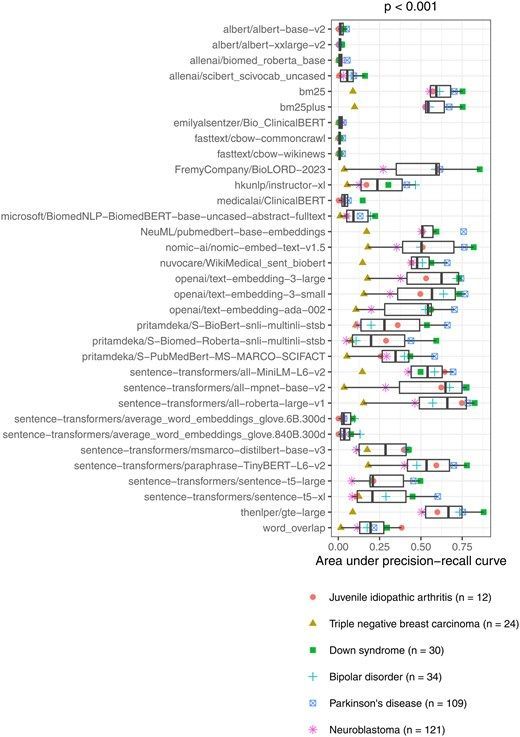
Using semantic search to find publicly available gene-expression datasets academic.oup.com/bioinformatics… 🧬💻🧪 github.com/srp33/GEO_NLP



English
Ben Busby! 리트윗함

Ben Busby! 리트윗함

Looking forward to discussing my perspectives of #spatialomics on precision medicine this afternoon.
UNMC Dept of Genetics, Cell Biology & Anatomy@UNMC_GCBA
GCBA Seminar Series announces Lana Garmire, PhD, University of Alabama-Birmingham, Department of Biomedical Informatics and Data Science presenting "Translating Single-cell and Spatial Omics into Precision Medicine." @UNMC_GCBA @UNMC_BISB
English
Ben Busby! 리트윗함

🧬 Hiring Postdocs at @astar_gis !
Help us decode cancer genomes. We need Computer Scientists and Computational Biologists to develop novel algorithms for de novo assembly of cancer genomes or to help us reconstruct them.
Experience in sequence alignment/assembly algorithms or de novo assembly of complex genomes required
Passionate about algorithmic puzzles with clinical impact? DM me.
Please RT! 🔄 #CompBio #Genomics #Algorithms #Postdoc

English
Ben Busby! 리트윗함

First test of partial epigenetic reprogramming to reverse aging for two eye conditions (glaucoma and NAION) gets FDA go ahead
@lifebiosciences Based on 2020 @Nature paper pubmed.ncbi.nlm.nih.gov/33268865/
endpoints.news/exclusive-fda-…

English
Ben Busby! 리트윗함

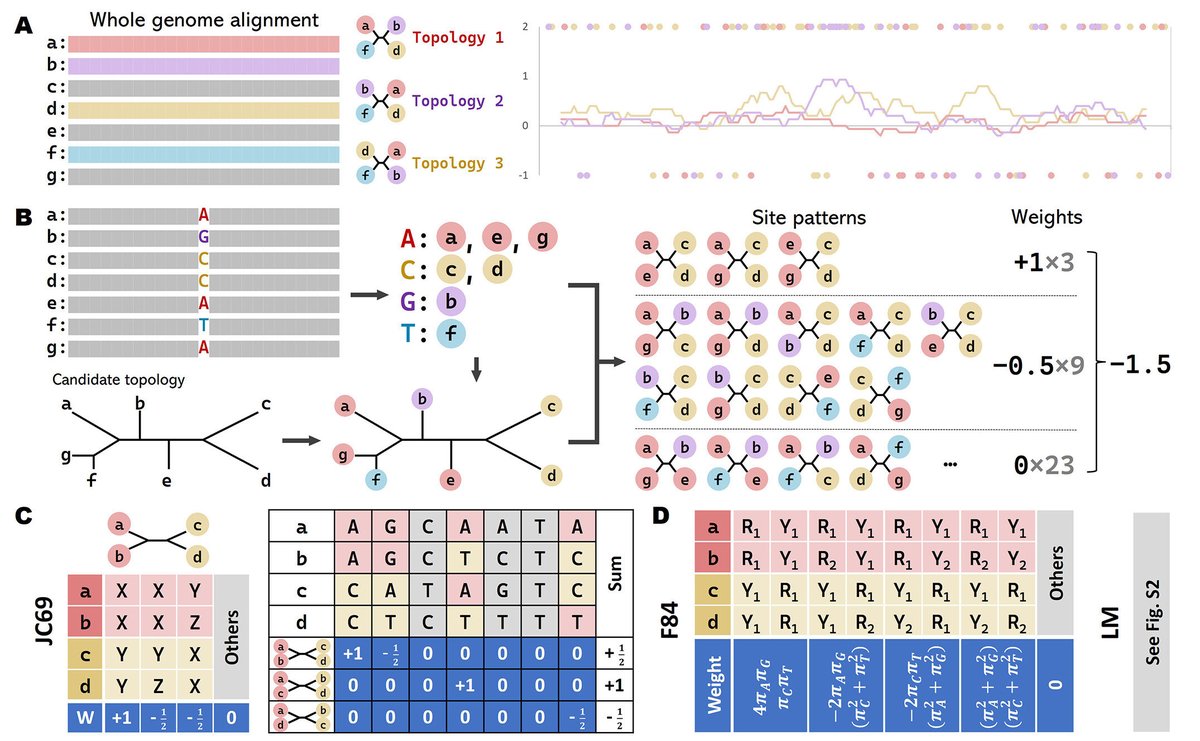
CASTER: Direct species tree inference from whole-genome alignments science.org/doi/10.1126/sc…
English
Ben Busby! 리트윗함

AlphaGenome: Decoding the dark matter of the genome with a unified deep learning model
More than 98% of human genetic variation lies outside protein-coding regions. These "non-coding" variants can disrupt gene regulation in remarkably diverse ways: altering chromatin accessibility, shifting 3D genome architecture, modifying splicing, or changing expression levels—often in tissue-specific patterns. Yet existing computational models face a fundamental trade-off: either they capture long-range regulatory interactions (like distant enhancers) but blur fine-scale features, or they achieve nucleotide resolution but miss distal context. And most specialize in a single modality, leaving users to stitch together predictions from many separate tools.
Žiga Avsec and coauthors at Google DeepMind present AlphaGenome, a model that sidesteps these trade-offs. It takes 1 megabase of DNA as input and predicts ~6,000 genome tracks—spanning gene expression, splicing (sites, usage, and junctions), chromatin accessibility, histone modifications, transcription factor binding, and 3D contact maps—at up to single-base-pair resolution.
The architecture combines a U-Net backbone with transformer blocks: convolutions capture local motifs essential for splice sites and TF footprints, while transformers model long-range dependencies like enhancer–promoter interactions. Training uses a two-stage approach—pretraining on experimental data followed by distillation from an ensemble of teachers using mutationally perturbed sequences—yielding a single model that scores variants across all modalities in one pass.
The results are striking: AlphaGenome achieves state-of-the-art performance on 25 of 26 variant effect prediction benchmarks, including a 25% improvement in predicting eQTL direction over the previous best model. It outperforms specialized models on their own tasks—beating SpliceAI-class methods on 6 of 7 splicing benchmarks and ChromBPNet on accessibility QTLs. Critically, the multimodal outputs enable mechanistic interpretation: for oncogenic mutations near the TAL1 gene in T-cell leukemia, AlphaGenome simultaneously predicts neo-enhancer formation (increased H3K27ac), chromatin opening, and elevated gene expression—recapitulating experimentally validated mechanisms.
This points toward a future where interpreting non-coding variation no longer requires assembling a patchwork of specialized models. A unified framework that jointly predicts molecular consequences across modalities could accelerate rare disease diagnostics, guide therapeutic oligonucleotide design, and help prioritize variants in GWAS loci—moving us closer to truly reading the regulatory code written in DNA.
Paper: nature.com/articles/s4158…

English
Ben Busby! 리트윗함

@ChelbyMustang researchers conducting research can hold them for SCIENCE
English