Hower Lee
42 posts

Hower Lee
@lee_hower
I work with spatial omic technologies and stuff. PhD student at Mats Nilsson Lab @matsnilssonlab


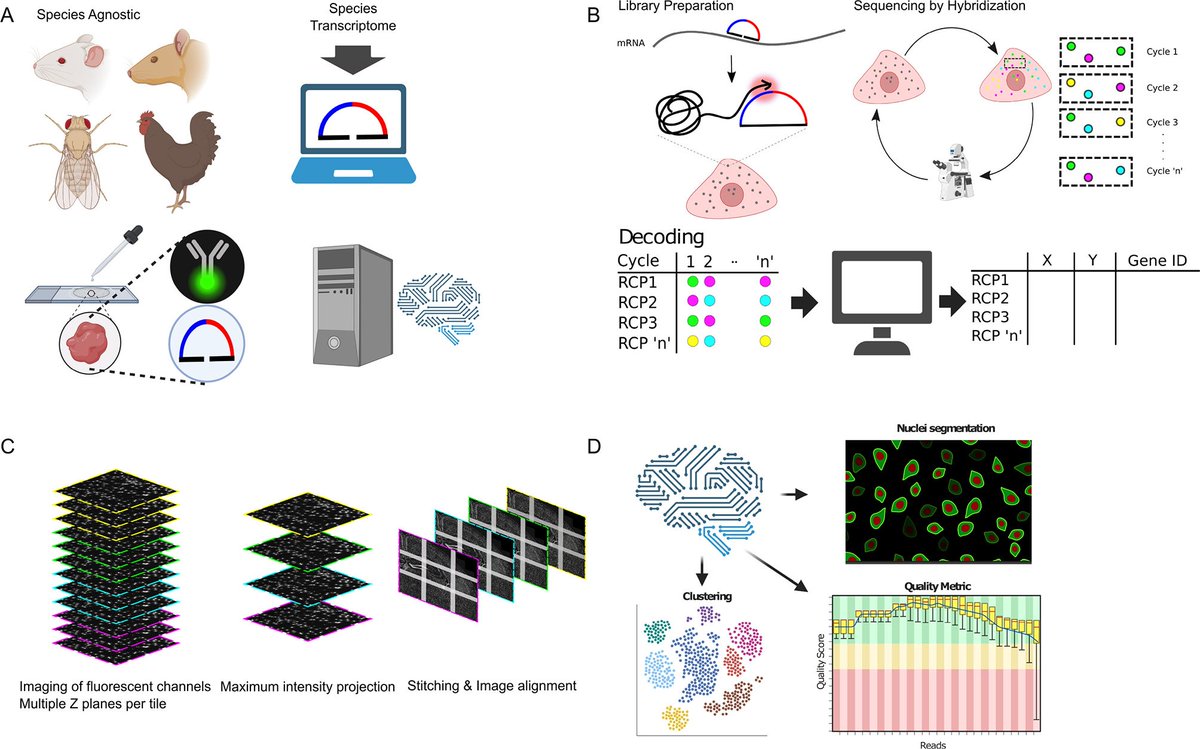




Open-source, high-throughput targeted in-situ transcriptomics for developmental biologists biorxiv.org/cgi/content/sh… #bioRxiv



🔥off the press, our new study with @MatsNilssonLab is online! Very proud of this work led by fearless and fabulous @IlonLiu, Li Jiang and Erik Samuelsson on the spatial organization of adult and pediatric H3K27M gliomas. @DFBC_PedCare @karolinskainst nature.com/articles/s4158…

Out now! Happy to share our @PLOSCompBiol publication w/ @CarolinaWahlby @gapartel on exploring the cellular architecture of the developing human #heart by applying segmentation-free algorithms #spage2vec on #nSituSequencing data. dx.plos.org/10.1371/journa…




Tgt w @joalunatkthse, @EMBO and Samakovlis Lab, we successfully concluded the first “Spatial Analysis of Gene Expression in Tissues” course @ @scilifelab last week. A huge thank you to all the course instructors and participants that made this event an overwhelming success 🤩



@dvd_bruggen's last piece of work in the lab, on the first waves of human oligodendrogenesis, is out @Dev_Cell! cell.com/developmental-… w/ Erik Sundström Fabio Pohl @PKukanja @mandy_meijer @kabbe_m @MossiAlejandro @slinnarsson @MatsNilssonLab @chrislangseth @M_Hilscher @lee_hower
