Sabitlenmiş Tweet

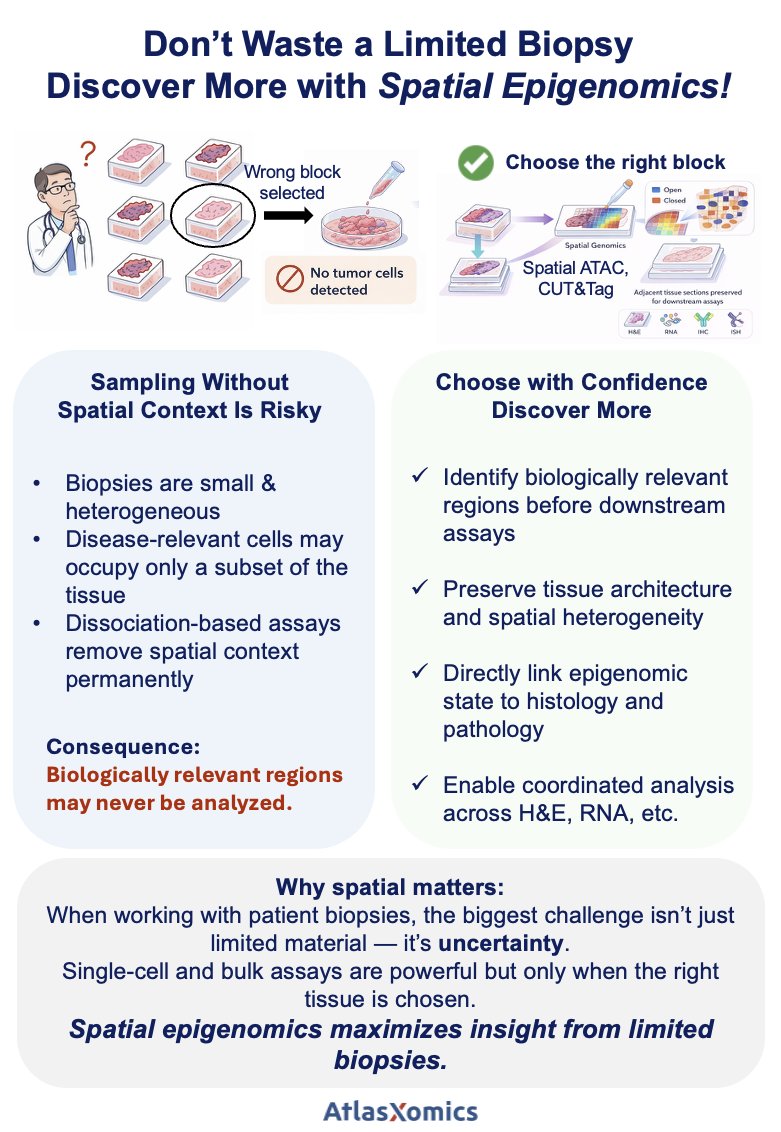
How tightly does chromatin accessibility reflect gene expression — in the same tissue?
With Spatial CoProfiling, we measure chromatin accessibility and whole-transcriptome RNA from the same tissue section, enabling direct, spatially resolved regulatory inference.
In our latest dataset, we observe:
Concordant accessibility and expression → active regulation
Accessibility preceding expression → poised states
Discordant signals → alternative regulatory mechanisms
This integrated view links chromatin state to transcription — in situ.
👉 Explore the dataset and white paper: atlasxomics.com/coprofiling-wh…
#SpatialEpigenomics #SpatialOmics #SpatialTranscriptomics #GeneRegulation

English