Bryce Kille
18 posts

Bryce Kille
@BKille
PhD student and former software engineer in the Treangen lab at Rice University. MS in computer science from UIUC.

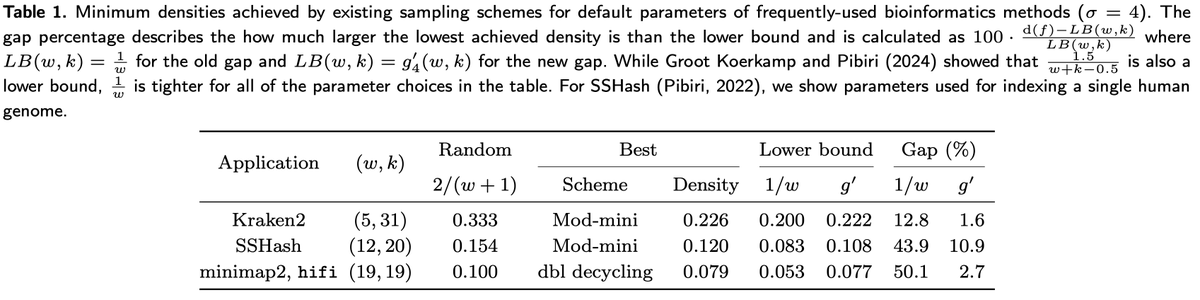
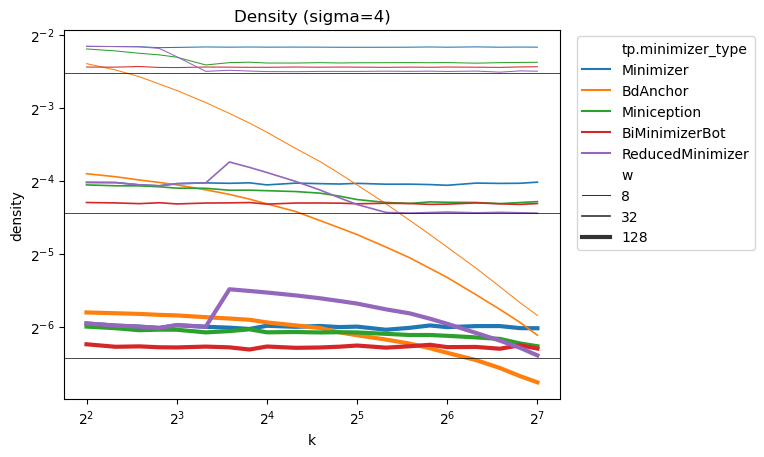
In a new preprint, @curious_coding and I led the derivation of a new lower bound, g’ (red), on k-mer sampling scheme density: - our bound appears to be tight for k ≡ 1 (mod w). - existing schemes are nearly optimal! - mod-minimizers are optimal for large sigma for k ≡ 1 (mod w)



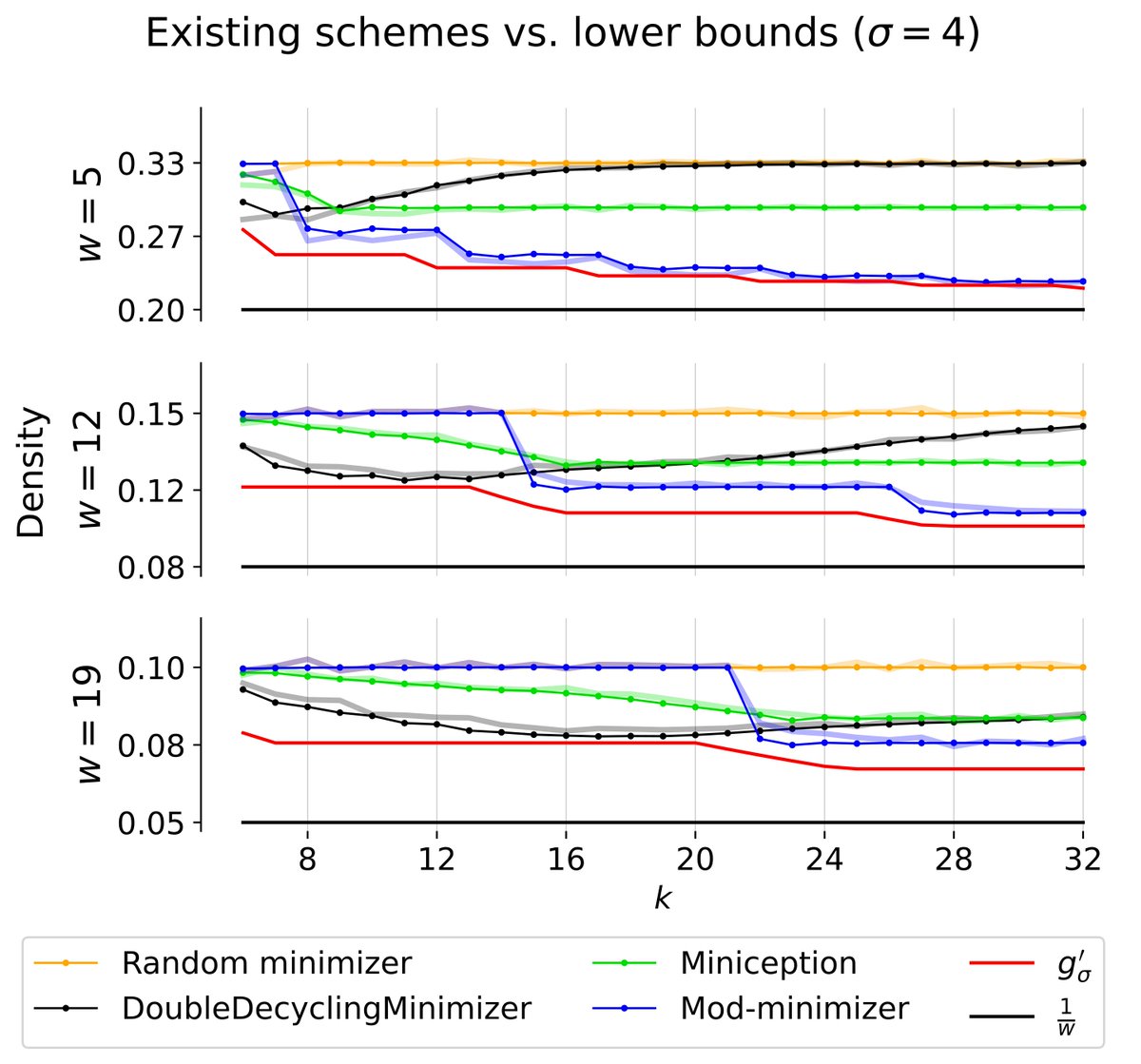
A near-tight lower bound on the density of forward sampling schemes biorxiv.org/cgi/content/sh… #biorxiv_bioinfo





For anybody using (l=31, k=21) minimizers, I have an exciting announcement coming up as soon as I finish writing stuff down. Saves 10% density using 2 if statements!




Our latest methods paper "Minmers are a generalization of minimizers that enable unbiased local Jaccard estimation" (aka MashMap3) is out with @BKille @erikgarrison @traingene! There's an interesting backstory here 🧵 \ biorxiv.org/content/10.110…






Rice CS PhD student Tiancheng Xu has developed a new hardware accelerator that speeds up genome variant analysis, processing #SARSCoV2 data in minutes. He recently presented this new tool at #FCCM22 alongside his advisors Alan Cox and Scott Rixner. bit.ly/3zNdypw

Very happy to share the latest paper from my group, SeqScreen, just published in @GenomeBiology. SeqScreen was led by my @RiceCompSci PhD students @AdvaitBalaji & @BKille, spanning 5 fruitful years of collaboration with Signature Science LLC @SigSci genomebiology.biomedcentral.com/articles/10.11… 1/n