And we're now published in Nature Methods 🥳 Full article here: nature.com/articles/s4159…
Geir Kjetil Sandve@SandveGeir
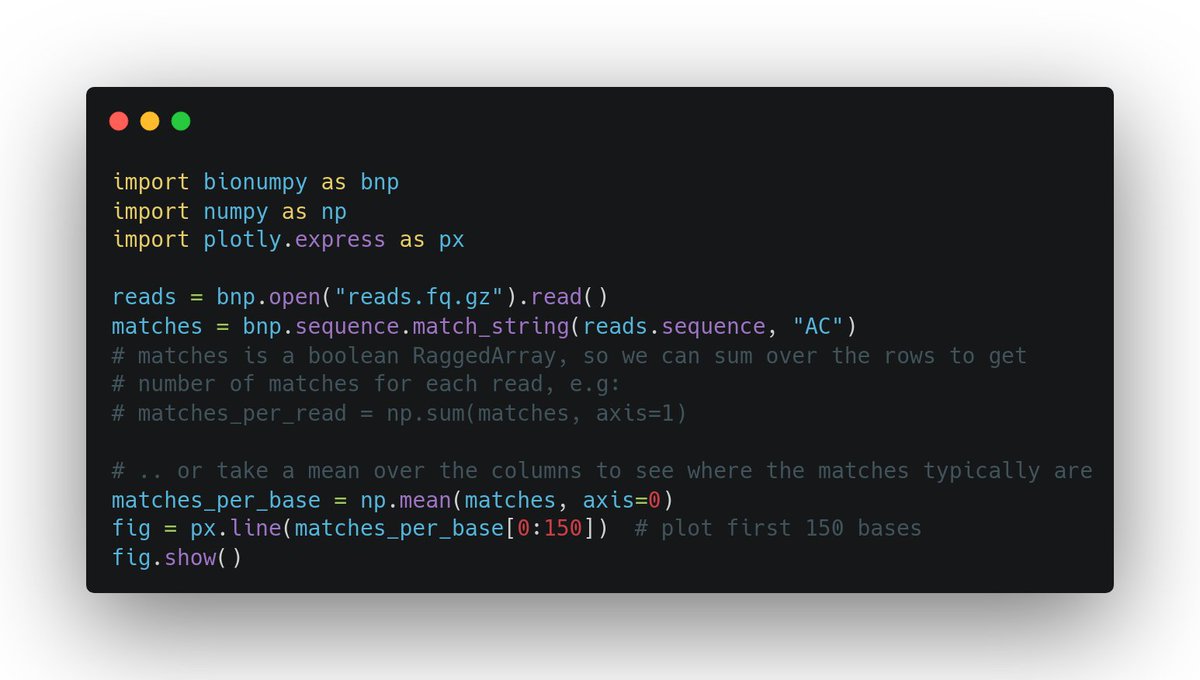
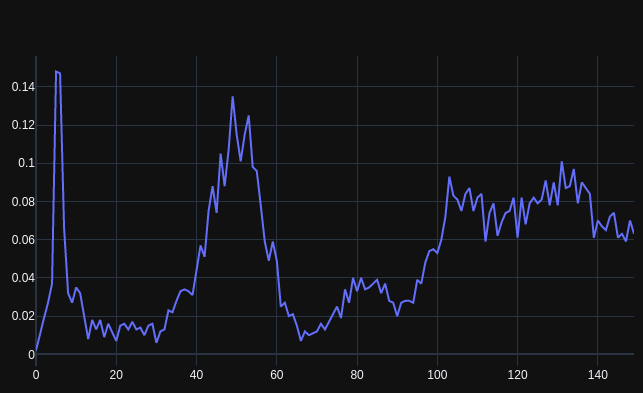
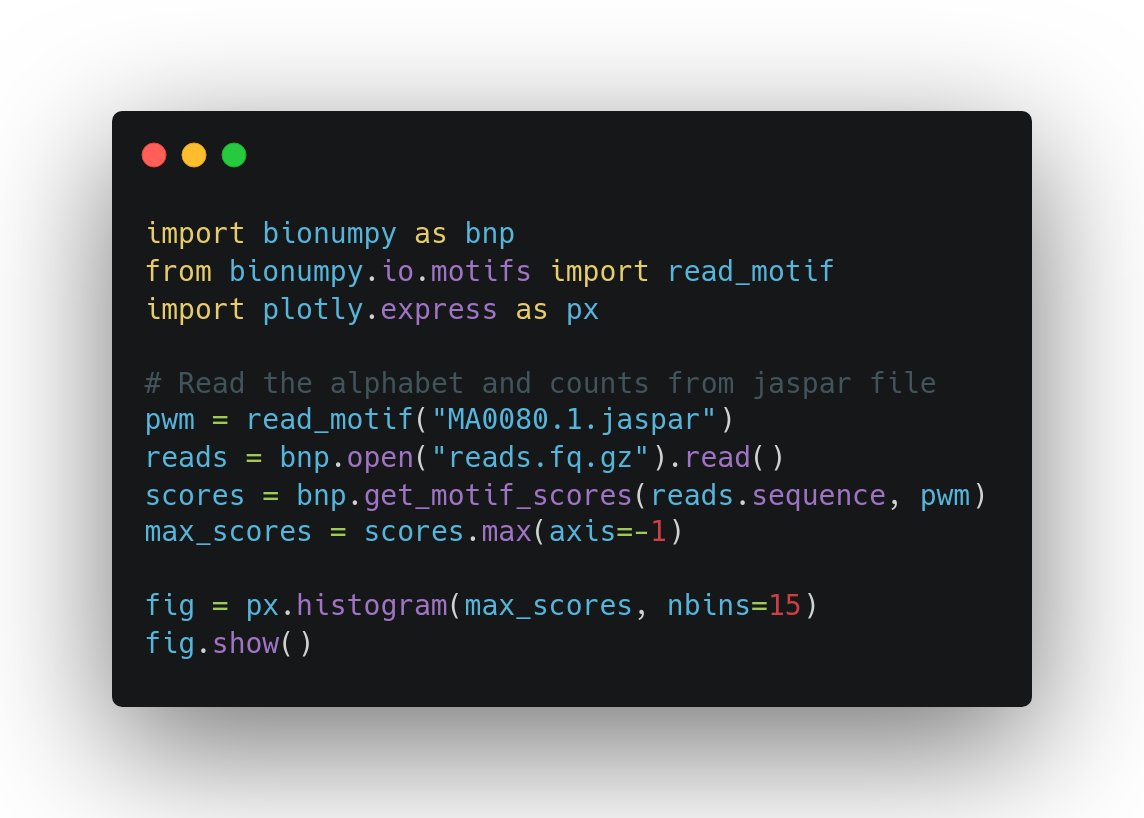
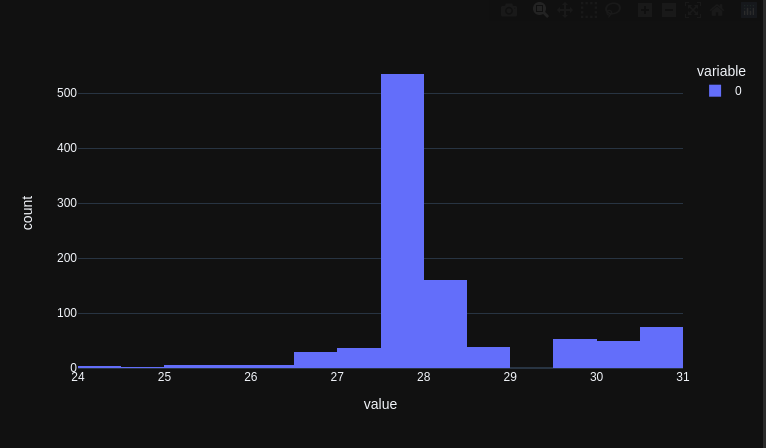
Finally biologists can also use numpy (array programming). Handling e.g. DNA and protein sequences with convenience and speed, like physicists and machine learners for decades have worked with numerical data: nature.com/articles/s4159… (1/3)
English