Daniel Portik
852 posts


Daniel Portik
@DPortik
Bioinformatics scientist @PacBio. I work on metagenomics, methylation, and assembly, but also enjoy phylogenomics and frog/lizard evolution. 🦠 🐸 🦎 🧬


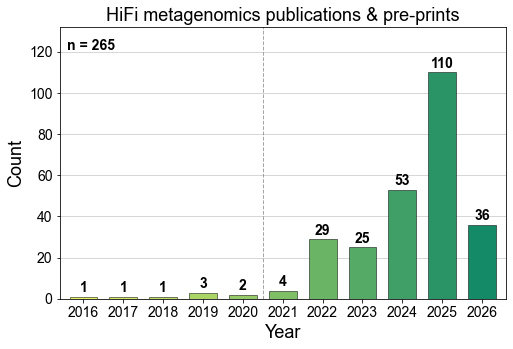























Preprint alert 🚨 Cautions in the use of @nanopore sequencing to map DNA modifications: officially reported “accuracy” ≠ reliable mapping in real applications. We performed a critical assessment of nanopore sequencing (across different versions of models) for the detection of multiple forms of DNA modifications: while it is reliable for high-abundance modifications, including 5mC at CpG sites in human cells and 5hmC in human brain cells, it makes very high % false positive calls for low-abundance modifications, such as 5mC at CpH sites, 5hmC and 6mA in most human cell types [1]. Want to know why? Please read further (long): In 2022, we reported in @ScienceMagazine that 6mA is much less abundant in human cells than previously reported (mostly false positives) [2]. In 2023, we wrote a perspective in @NatureRevGenet to help navigate the common pitfalls for false positive in the mapping of DNA and RNA modifications [3]. With these two papers, we thought the importance of recognizing false positive calls has been adequately described. Unfortunately, this was not the case. New official models report nanopore seq can achieve >97.5% raw read accuracy in the detection of 5mC, 5hmC and 6mA, across all sequence contexts. New methods are to be released for mapping DNA damages, etc. Unfortunately, we do not think these “accuracies” are reliable in practical applications. To help biologists and methods developers, @kong_yimeng (first author of the 2022 & 2023 papers, now a PI) and I felt the need to do this study to critically assess nanopore seq for the detection of multiple forms of DNA mods. Thanks to team and collaborators: @KohliLab @ChristianLoo_ @XuesongRutgers. Why did we pick nanopore in this critical assessment? First of all, we have no bias among different sequencing platforms. In fact, we regularly use nanopore seq in our lab for several projects. However, we think there is a broad misunderstanding and over expectation that nanopore sequencing can map many forms of DNA modifications in any genome. A fundamental limitation is that the training and test data used in existing machine learning models do not consider the physiological levels of DNA modifications. Common pitfalls: 1. High accuracies in training and test data don't guarantee reliability in real applications. It depends on the abundance of DNA modifications (6mA is abundant in bacteria, but rare in human). 2. Reliability in one sequence context or genomic region doesn't ensure reliability in others. It depends on the abundance of DNA modifications at a specific sequence context or genomic region (5mC is abundant at CpG sites but not at CpH sites; 5mC is abundant on the nuclear genome but not on mtDNA). 3. Reliability in one cell type doesn't guarantee success in others. It depends on the abundance of DNA modifications in a specific cell type (5hmC is abundant in brain tissues but rare in blood). Why do each of these pitfalls involve the critical consideration of the abundance of DNA modification? This is because every genomic technology has a certain level of background noise in detecting DNA modifications. It will detect a certain level of DNA modification even when no modification was present in a negative control sample! Therefore, the method is only reliable if the physiological levels of DNA modification are sufficiently higher than background noise. How to navigate the pitfalls? To assist biologists and methods developers, we describe a framework for rigorous evaluation that highlights the use of false discovery rates (FDRs) along with rationally designed negative controls capturing both general background and confounding modifications. What is “confounding modification”? It turns out that modification-free negative controls are not enough. For example, the abundant 5mCpG in most human cell types can lead to increased levels of false positive 5hmC calls, much higher than the actual 5hmC levels in most human cell types (Fig. 1 attached) …… (1) This study highlights the urgent need to incorporate this framework in future methods development and biological studies. Using this framework, you can quantitatively evaluate the reliability of DNA modifications detected in your samples. (2) This study advocates prioritizing nanopore sequencing for mapping abundant, not rare, modifications in biomedical applications. Specifically, 5mCpG can be largely reliably mapped across mammalian cell types; 5hmCpG mapping by existing methods should be limited to tissues with high 5hmC abundance (e.g. brain tissues). In addition, our assessment does not support the use of nanopore sequencing to map endogenous* 6mA or various DNA damages in mammalian genomes due to their low abundance. (3) This framework is also applicable to RNA modifications. Although this work is focused on DNA modifications, the principles we outline—rigorous false positive recognition, use of negative controls, and FDR evaluation—are equally applicable to RNA modifications. [1] New preprint: biorxiv.org/content/10.110… [2] Critical assessment of DNA adenine methylation in eukaryotes using quantitative deconvolution science.org/doi/10.1126/sc… [3] Navigating the pitfalls of mapping DNA and RNA modifications nature.com/articles/s4157… [*] Please note the detection of exogeneous DNA modifications is different from mapping natural endogenous DNA modifications, because abundant exogeneous DNA modifications are added to the genome as in Fiber-seq, DiMeLo-seq, etc.



