Sabitlenmiş Tweet

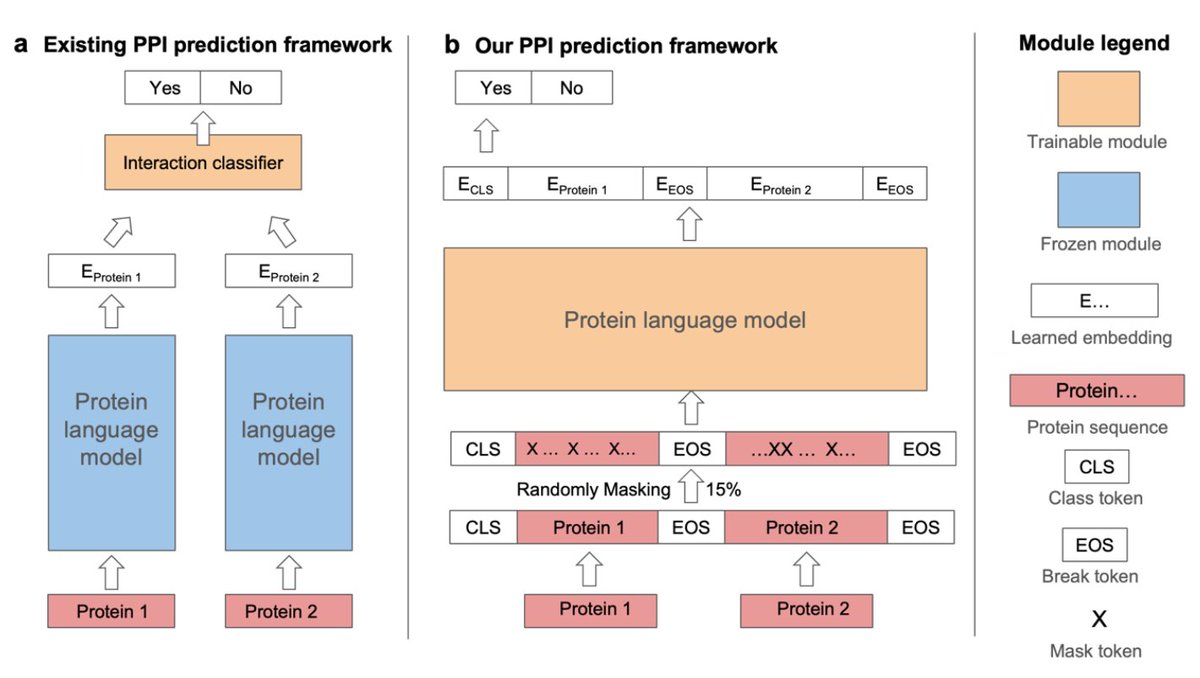
Our PLM-interact is out in @NatureComms! We show that jointly encoding protein pairs using protein language models improves protein–protein interaction prediction performance and enables fine-tuning to predict mutation effects in human PPIs.
nature.com/articles/s4146…
English