
Arthur Declercq
23 posts


@DeclercqArthur
PhD student at @CompOmics (VIB-UGent), @VIBLifeSciences Bioinformatics | Machine learning | Immunopeptidomics













It really is free and less than 2 weeks away. Register now for the EuPA and EPIC-XS workshop and webinar on Computational Proteomics compomics.com/events/2022-ep…





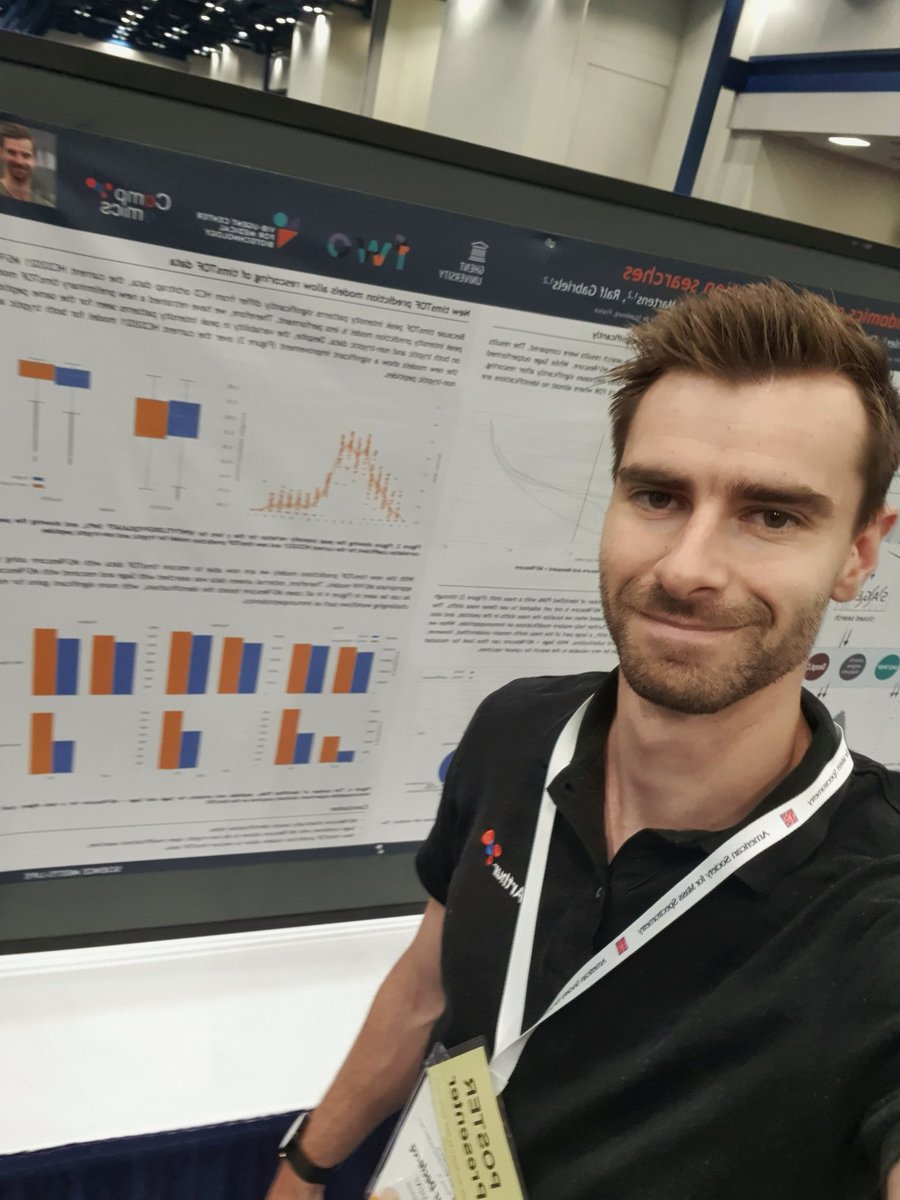
Here's an overview of @CompOmics at #HUPOReCONNECT 2021. Checkout out posters today at 16:00 UTC, Sven's talk on #ionbot Friday at 0:55 UTC, and the session "The Future of Proteomics" Lennart is chairing with @mlaval6, also on Friday.



Manuscript on our top performing immunopeptidomics identification engine now out in bioRxiv: biorxiv.org/content/10.110… - if you are interested in 30% more unique identified immunopeptides, or if you care about top performance even at 0.1% FDR (!!), then definitely check it out!

Manuscript on our top performing immunopeptidomics identification engine now out in bioRxiv: biorxiv.org/content/10.110… - if you are interested in 30% more unique identified immunopeptides, or if you care about top performance even at 0.1% FDR (!!), then definitely check it out!

Manuscript on our top performing immunopeptidomics identification engine now out in bioRxiv: biorxiv.org/content/10.110… - if you are interested in 30% more unique identified immunopeptides, or if you care about top performance even at 0.1% FDR (!!), then definitely check it out!