Vadim Demichev
1.5K posts

Vadim Demichev
@DemichevLab
Group Leader at @ChariteBerlin. Proteomics technologies and applications, DIA-NN author.



Software horror: litellm PyPI supply chain attack. Simple `pip install litellm` was enough to exfiltrate SSH keys, AWS/GCP/Azure creds, Kubernetes configs, git credentials, env vars (all your API keys), shell history, crypto wallets, SSL private keys, CI/CD secrets, database passwords. LiteLLM itself has 97 million downloads per month which is already terrible, but much worse, the contagion spreads to any project that depends on litellm. For example, if you did `pip install dspy` (which depended on litellm>=1.64.0), you'd also be pwnd. Same for any other large project that depended on litellm. Afaict the poisoned version was up for only less than ~1 hour. The attack had a bug which led to its discovery - Callum McMahon was using an MCP plugin inside Cursor that pulled in litellm as a transitive dependency. When litellm 1.82.8 installed, their machine ran out of RAM and crashed. So if the attacker didn't vibe code this attack it could have been undetected for many days or weeks. Supply chain attacks like this are basically the scariest thing imaginable in modern software. Every time you install any depedency you could be pulling in a poisoned package anywhere deep inside its entire depedency tree. This is especially risky with large projects that might have lots and lots of dependencies. The credentials that do get stolen in each attack can then be used to take over more accounts and compromise more packages. Classical software engineering would have you believe that dependencies are good (we're building pyramids from bricks), but imo this has to be re-evaluated, and it's why I've been so growingly averse to them, preferring to use LLMs to "yoink" functionality when it's simple enough and possible.

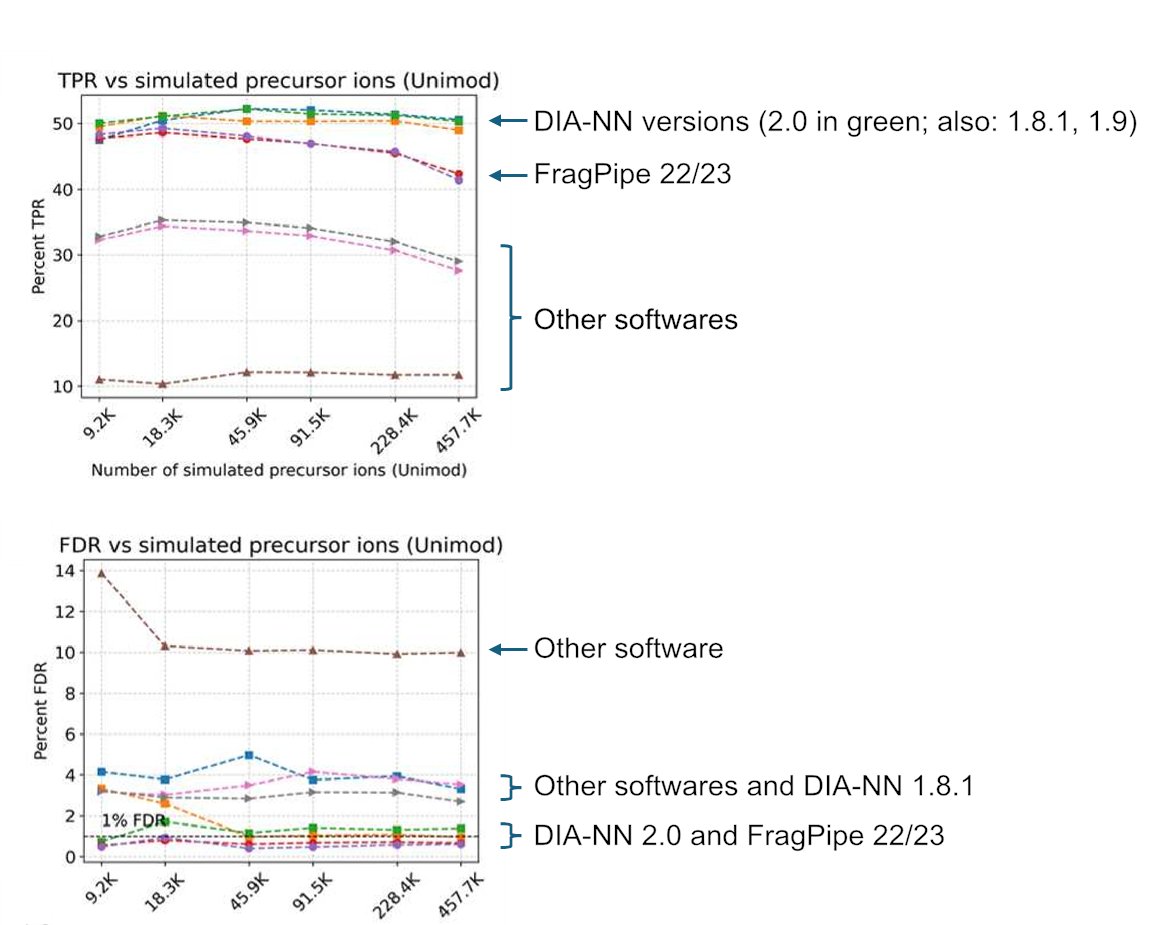
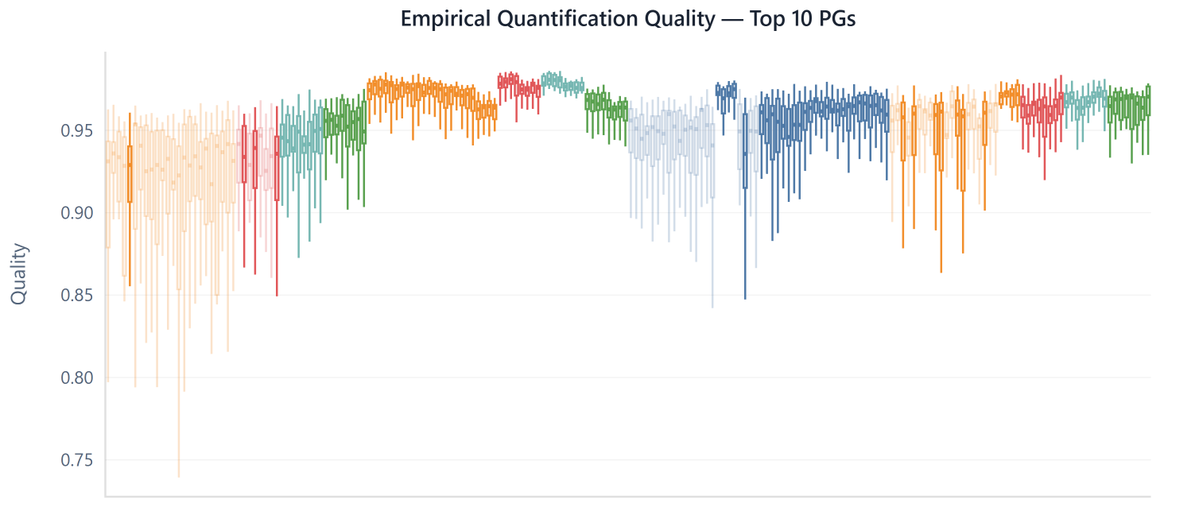
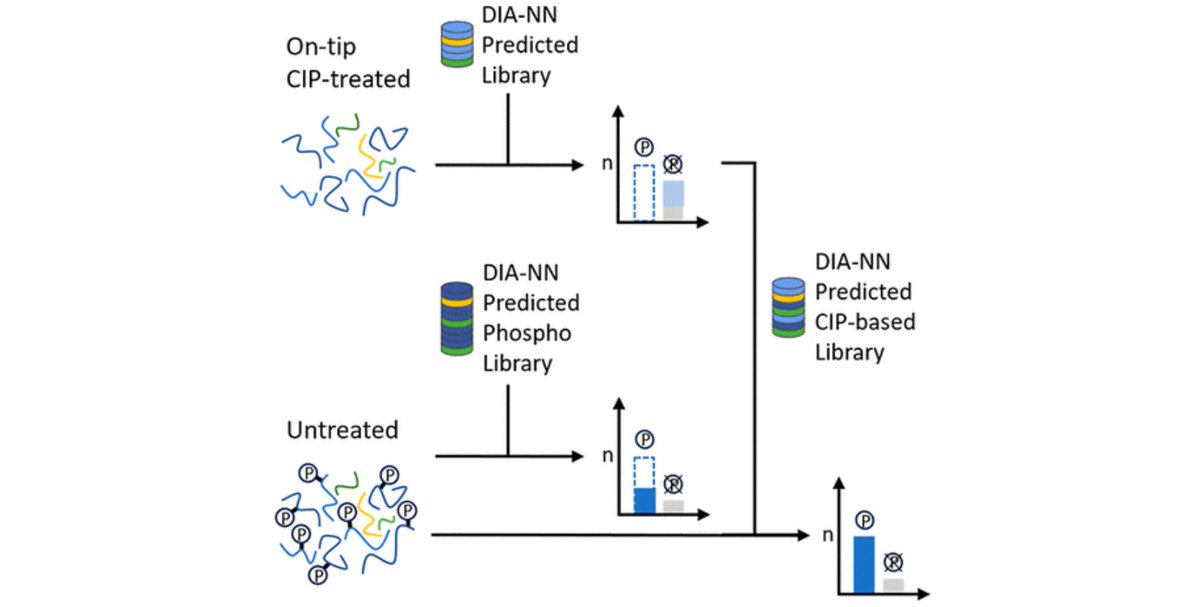
Our benchmarks reveal inflated true FDR (3–5%) when modified peptidoforms are considered and highlight peak-matching errors in match-between-runs analyses. Great collaboration with the labs of Andreas Hildebrandt and Mathias Wilhelm. (2/2) Preprint: researchsquare.com/article/rs-903…