Francisco Gimeno-Valiente
46 posts


Francisco Gimeno-Valiente
@F_gimenoval
PhD in Biomedicine & Biotechnology | Cancer Researcher at UCL
London - UK Katılım Mayıs 2024
92 Takip Edilen48 Takipçiler

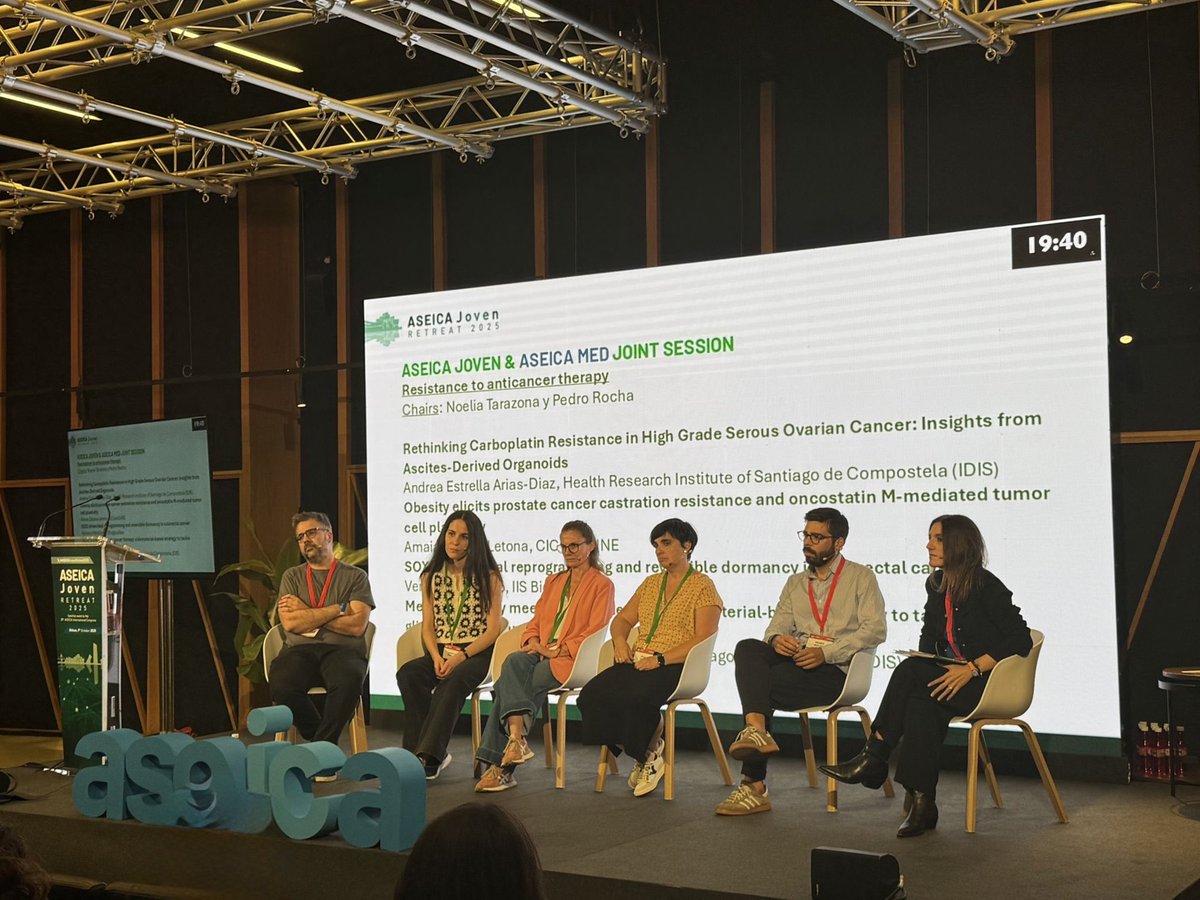
Amazing session at @ASEICAnews 2025 on resistance to anticancer therapy, chaired by @TarazonaNoelia and @pedrofsrocha 🔬🧬

English
Francisco Gimeno-Valiente retweetledi

🚨The SERENA-6 trial shows that acting before radiologic progression can matter.
Switching to camizestrant upon emerging ESR1 mutations in ctDNA🩸extended PFS and delayed QoL decline in HR+/HER2-BC.
👟A step closer to real-time, biology-driven care. ✍️@OrtegaTolosa @evaciruelos
English
Francisco Gimeno-Valiente retweetledi

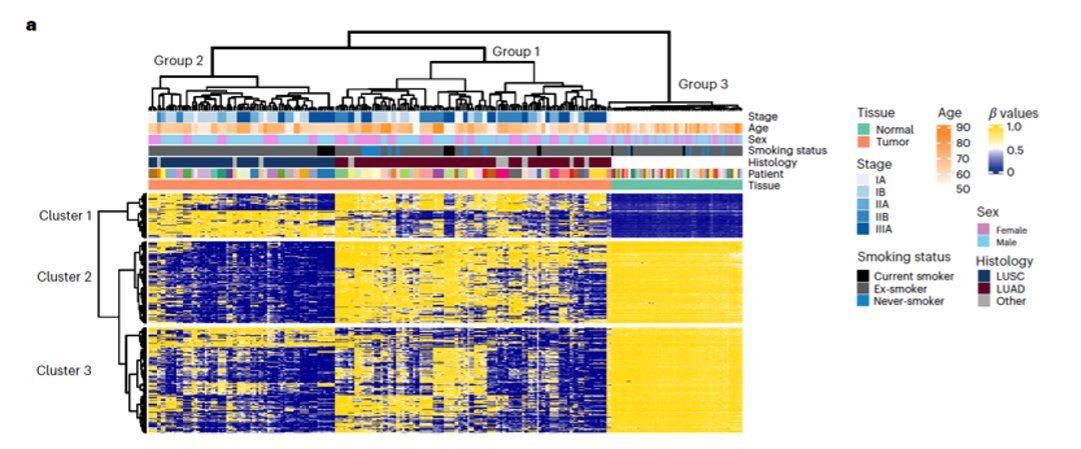
Great lecture by Nnenna Kannu @ucl at @myESMO MAP2025 featuring the DNA #methylation patterns of the amazing #lungcancer TRACERx cohort, now at @NatureGenet ! Congrats to @CharlesSwanton and @MariamJHanjani, too!

English
Francisco Gimeno-Valiente retweetledi

🚀 Hoy hemos celebrado la reunión de inicio del proyecto #MOLECULAR, dirigido a mejorar el abordaje del cáncer de cabeza y cuello localmente avanzado mediante el análisis del ADN tumoral en sangre e IA.
👩🔬 Esta iniciativa colaborativa está coordinada por la Dra. Gema Bruixola.

Español
Francisco Gimeno-Valiente retweetledi

(13/13)
Thank you to the patients and their families, without which we would not be able to start this project. Thank you to all our funders without which our work would not be possible @thecrick @crukcolcentre @cr_UK @uclcancer @rosetreest
English
Francisco Gimeno-Valiente retweetledi
Francisco Gimeno-Valiente retweetledi
Francisco Gimeno-Valiente retweetledi
Francisco Gimeno-Valiente retweetledi
Francisco Gimeno-Valiente retweetledi
Francisco Gimeno-Valiente retweetledi
Francisco Gimeno-Valiente retweetledi
Francisco Gimeno-Valiente retweetledi
Francisco Gimeno-Valiente retweetledi
Francisco Gimeno-Valiente retweetledi
Francisco Gimeno-Valiente retweetledi
Francisco Gimeno-Valiente retweetledi

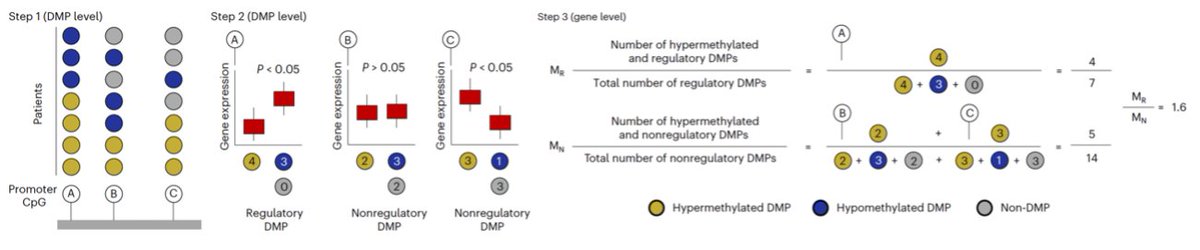
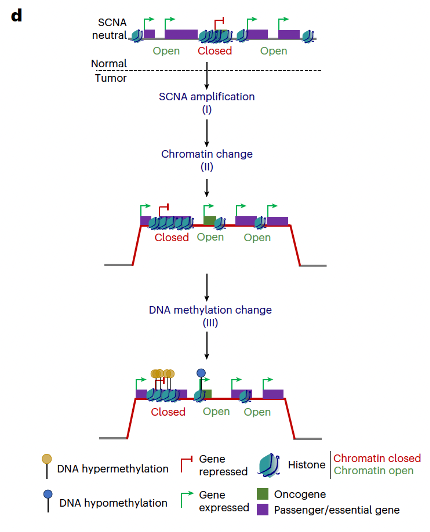
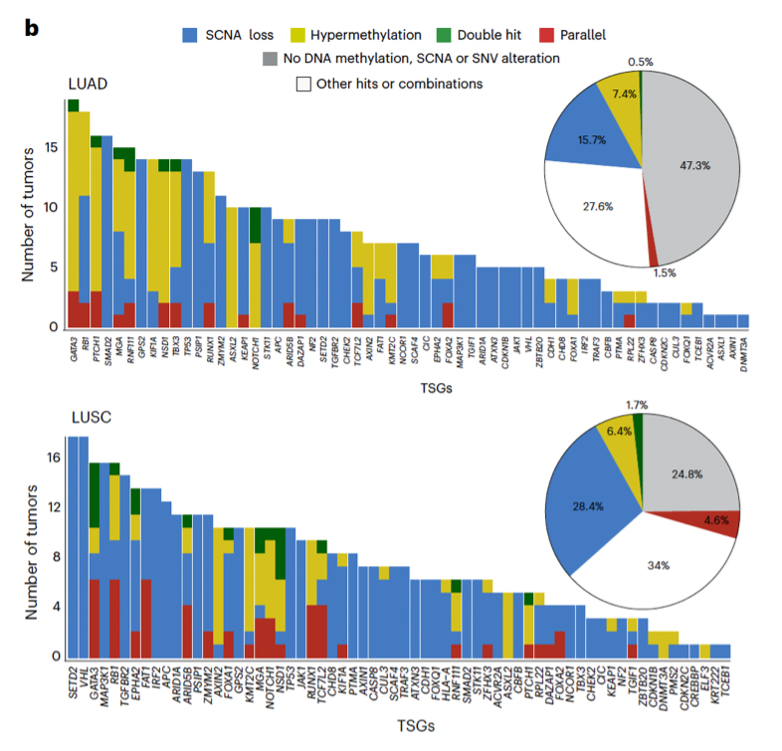
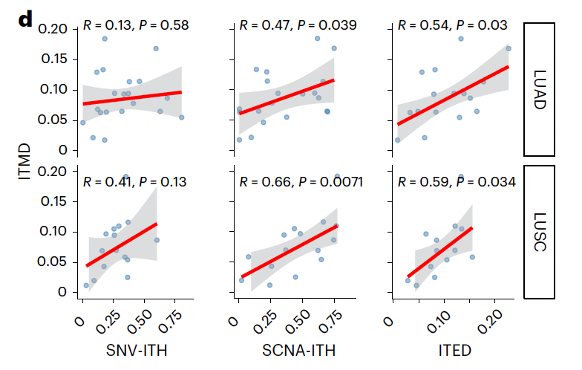
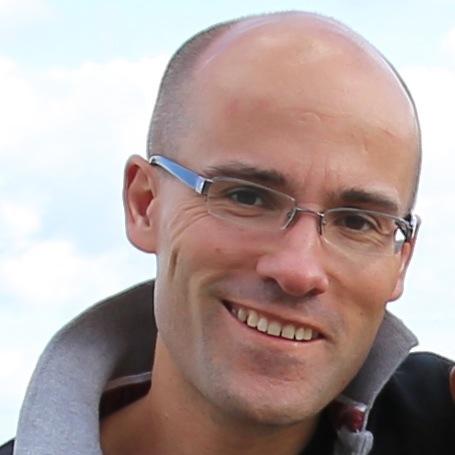
Proud to share our first TRACERx cancer study integrating epigenomics + genomics in NSCLC as co-drivers of tumor evolution.
Led by not me but by the wonderful @NnennayaKanu, @F_gimenoval, @ccastignani6, @VanLooLab


English
Francisco Gimeno-Valiente retweetledi
Francisco Gimeno-Valiente retweetledi

🔥New in @NatureGenet by @F_gimenoval during his postdoc at @ucl under @NnennayaKanu @CharlesSwanton 🔝
📌Genomic & epigenomic drivers converge in lung cancer. New methylation dN/dS + AllChAT reveal chromatin-driven dosage compensation.
🔗nature.com/articles/s4158…
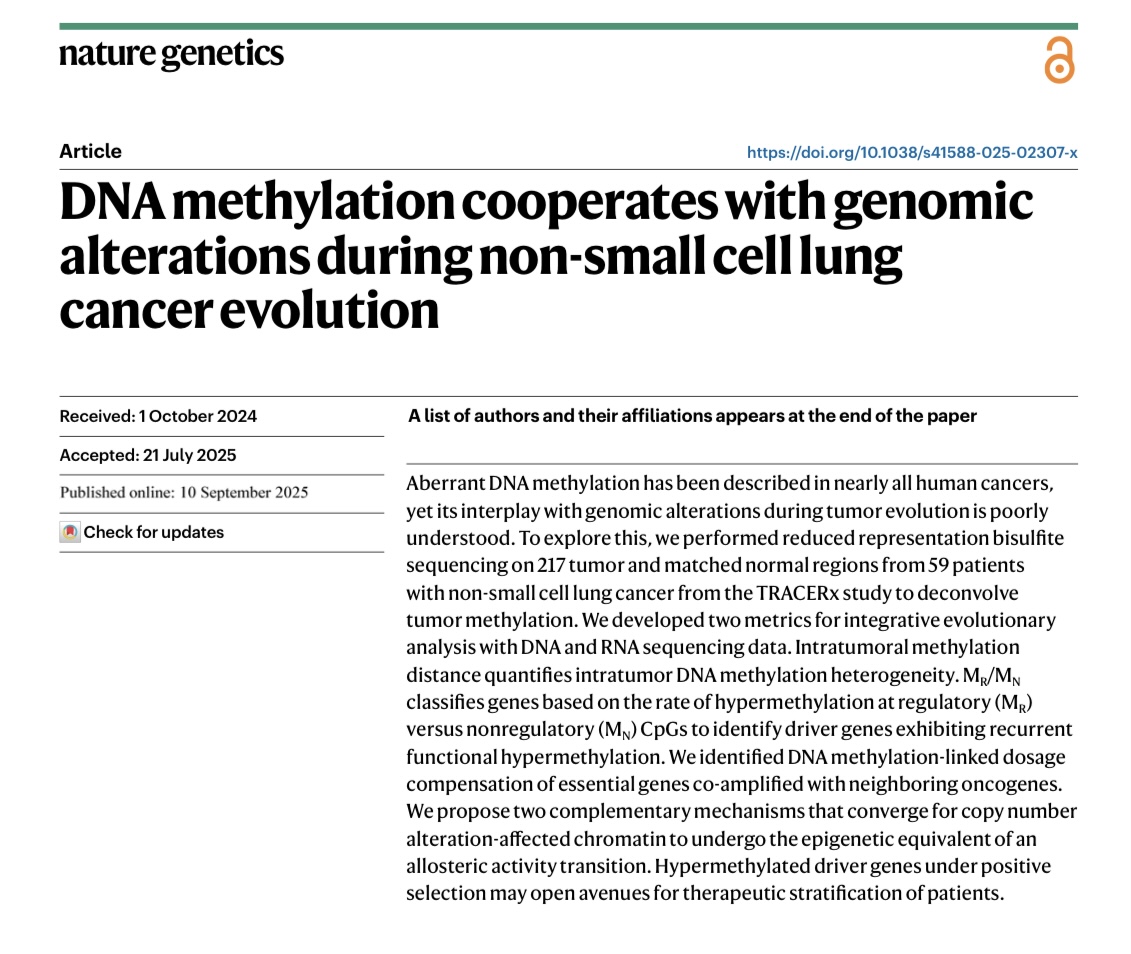


English