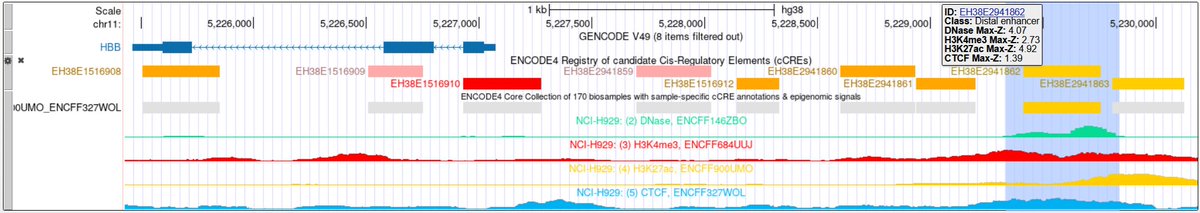
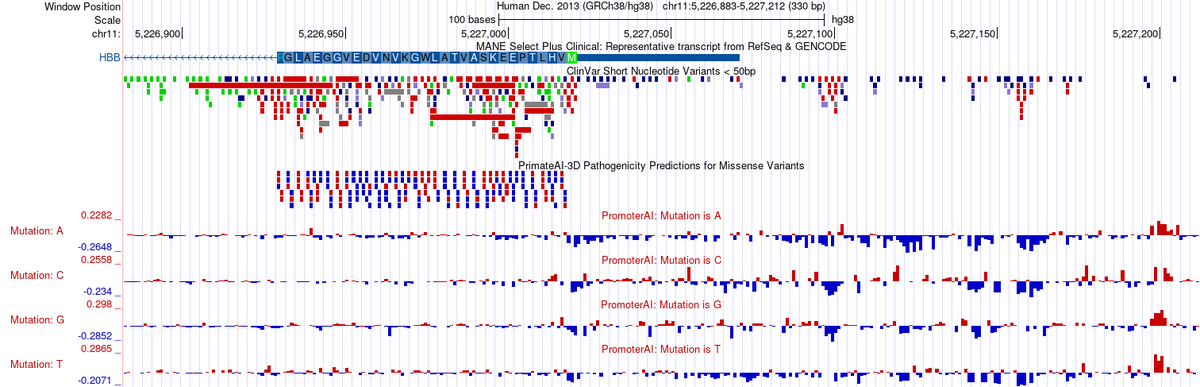
Two new variant-impact tracks from Illumina on the UCSC Genome Browser: PrimateAI-3D scores every coding missense variant (hg38/hg19); PromoterAI scores every non-coding substitution near transcription start sites (hg38). See our news for more: bit.ly/illuminaTracks
English