Jacob Loupe
84 posts


Jacob Loupe
@JLoupe2
Senior scientist in Myers lab @HudsonAlpha



So excited that our paper “De novo variants in the RNU4-2 snRNA cause a frequent neurodevelopmental syndrome” is out today in @Nature nature.com/articles/s4158… 🧵 1/16









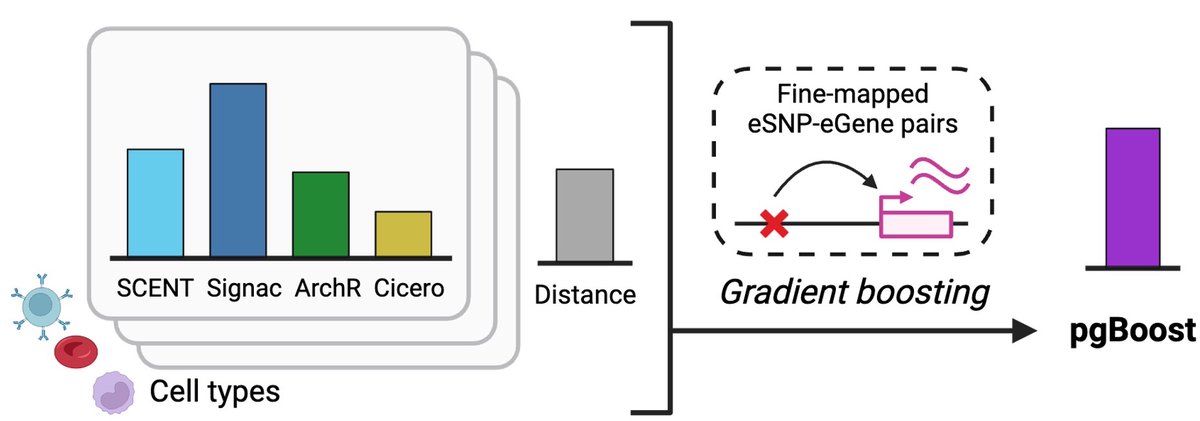
Linking regulatory variants to target genes by integrating single-cell multiome methods and genomic distance medrxiv.org/cgi/content/sh… #medRxiv
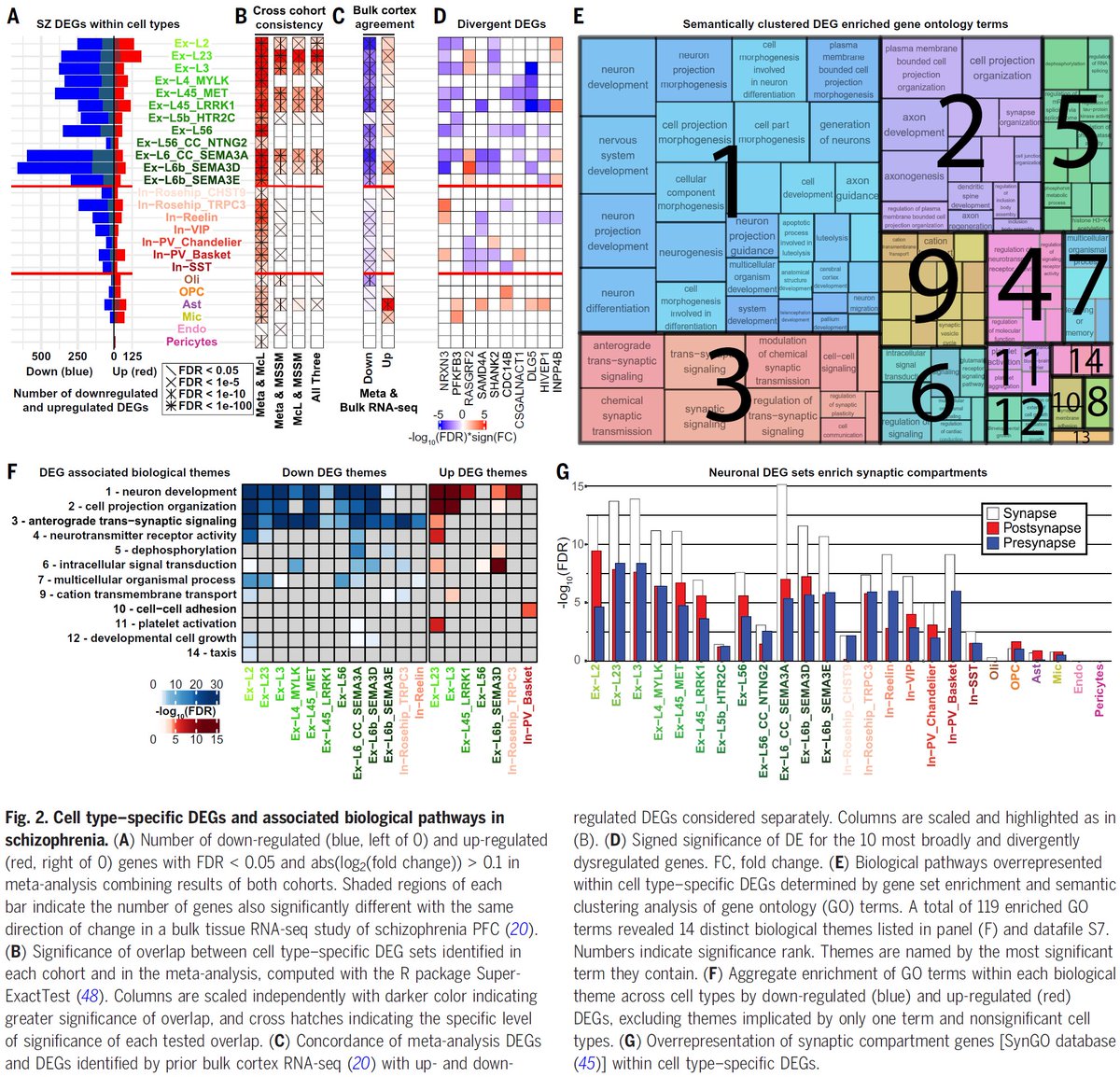



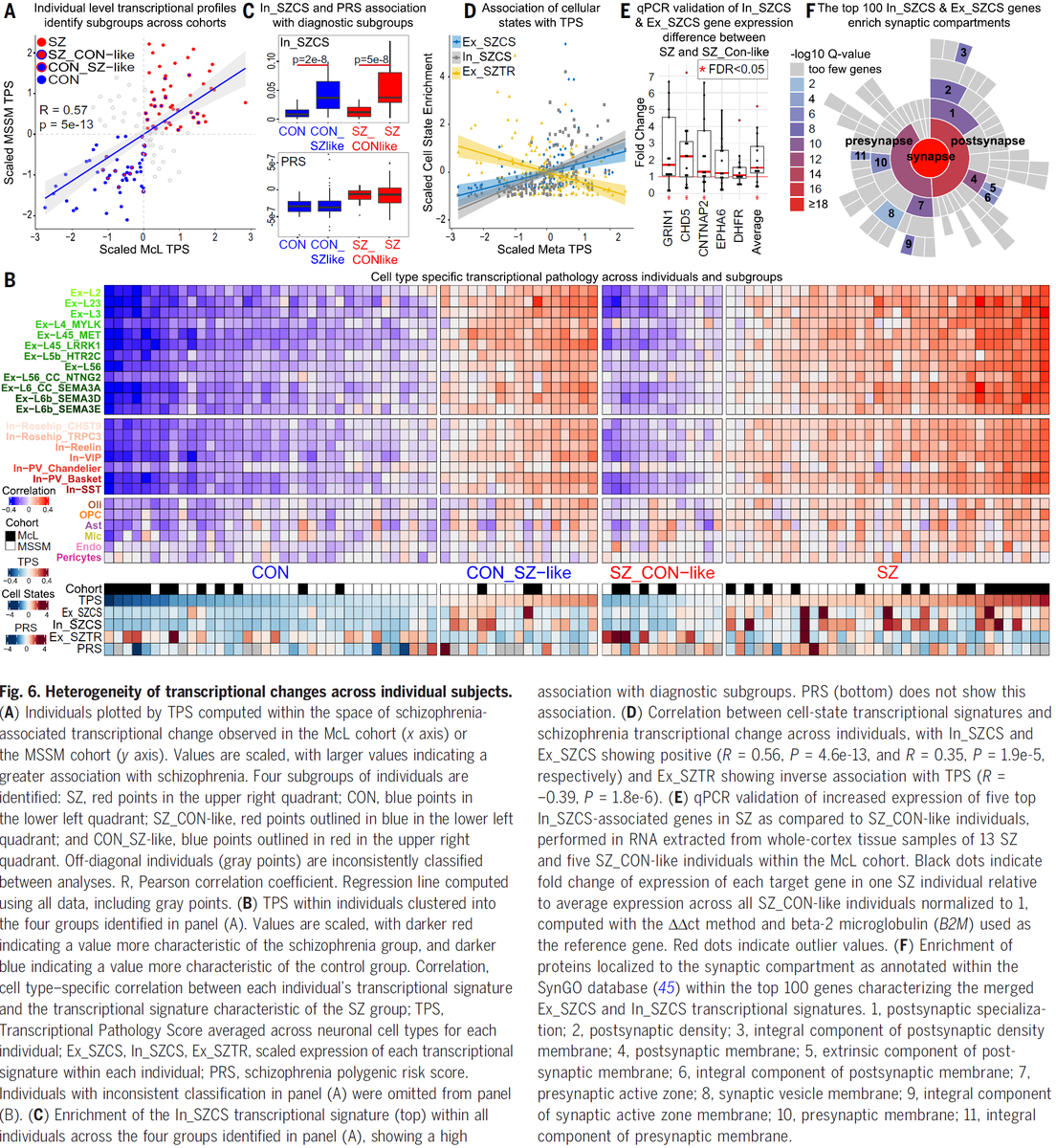
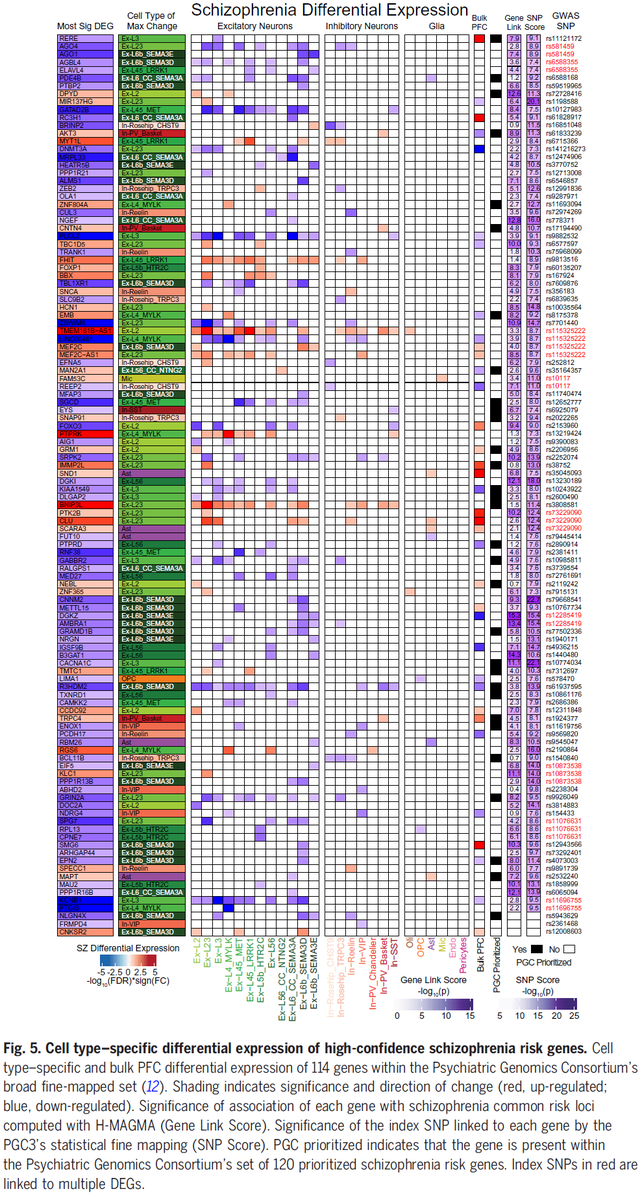
Our #SingleCell dissection of #Schizophrenia genes/pathways, a new #TranscriptionalResilience cell type, target genes at #GWAS loci, and upstream #MasterRegulators is out @medRxivPreprint #PsychENCODE #BradRuzicka @Mohammadi_PhD #JoseDavila @NIMHgov medrxiv.org/content/10.110…

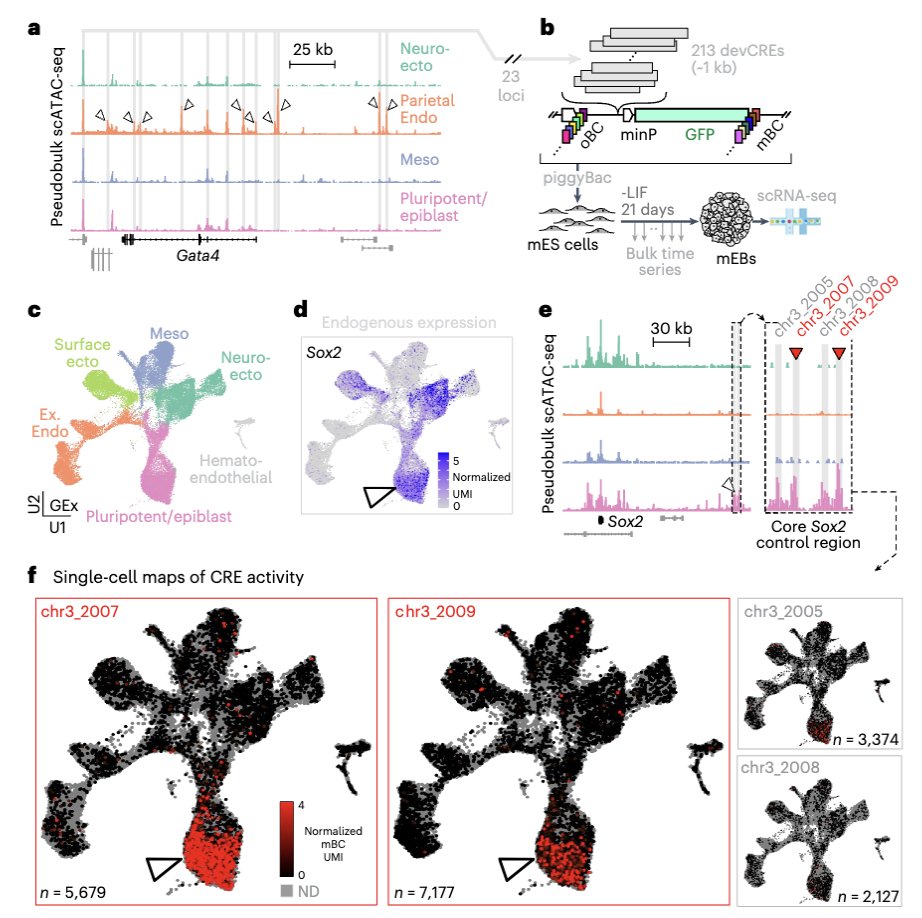



Multi-omic QTL mapping in early developmental tissues reveals phenotypic and temporal complexity of regulatory variants underlying GWAS loci biorxiv.org/cgi/content/sh… #bioRxiv

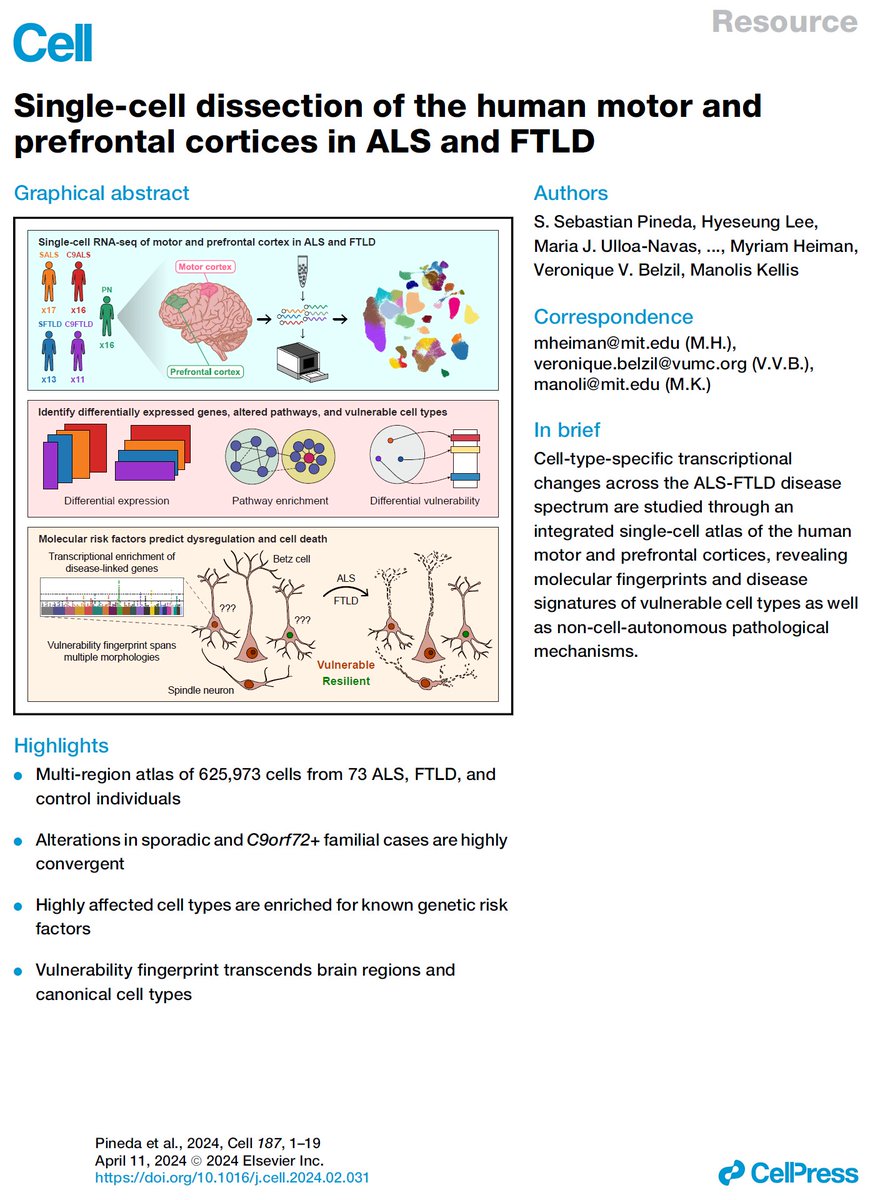
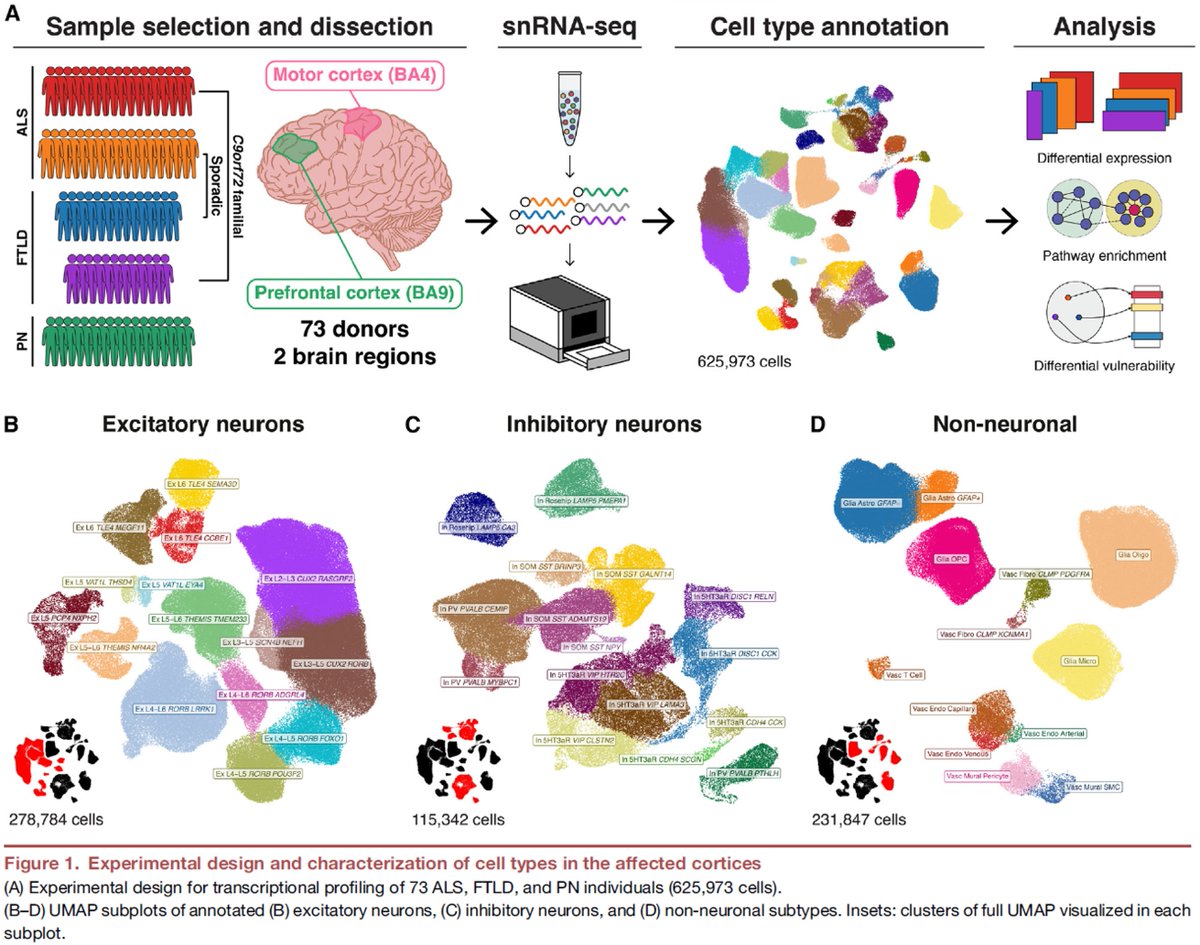


Excited to share our #SingleCell dissection of #MotorCortex in #ALS and #FTLD biorxiv.org/content/10.110… #BetzCells #VENs #C9orf72 #TDP43 #POU3F1 #ADRD #NeuroDegen #MCx #Alzheimer with #SebastianPineda #VeroniqueBelzil #MyriamHeiman #HyeseungLee @B_Fitzwalter @Mohammadi_PhD @MTPA_US