Sabitlenmiş Tweet

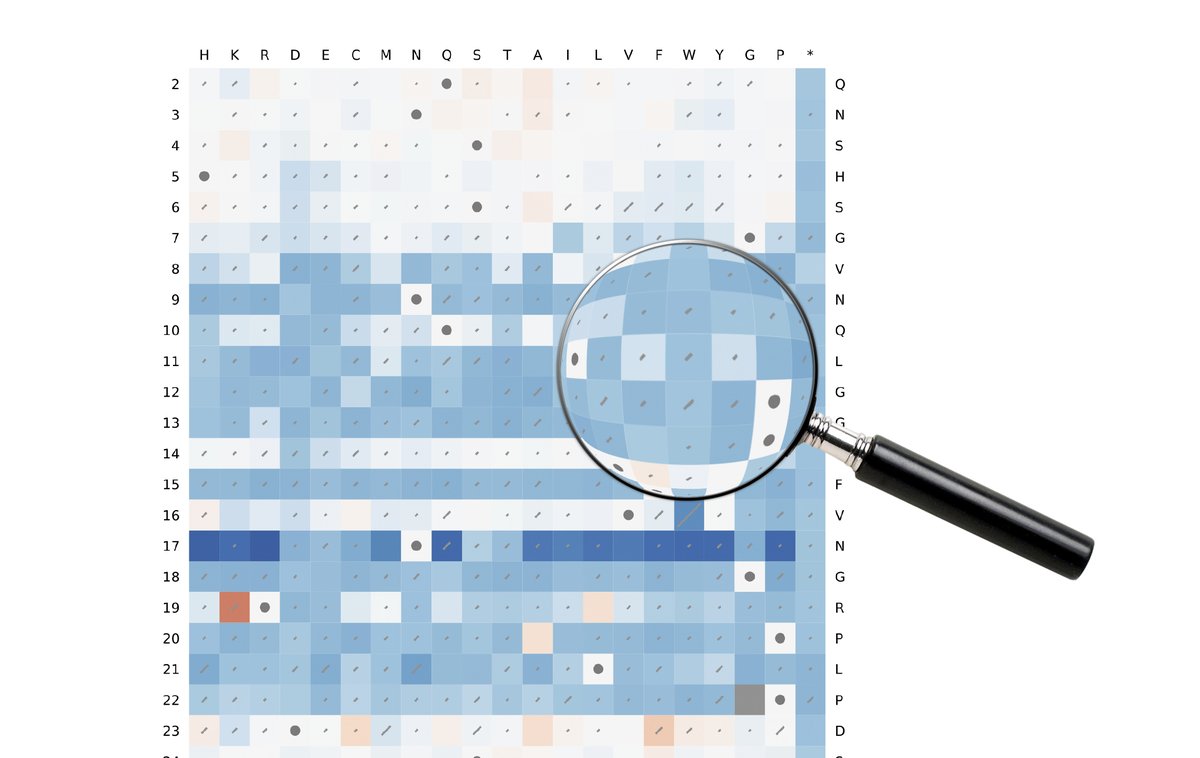
Our @jmarshlab/@MRC_HGU investigation of structural protein stability predictors using deep mutational scanning (DMS) data has been published in Protein Science @ProteinSociety doi.org/10.1002/pro.46…
English
Lukas Gerasimavičius
51 posts


@L_Geras
Research Fellow @jmarshlab | Institute of Genetics and Cancer, Edinburgh










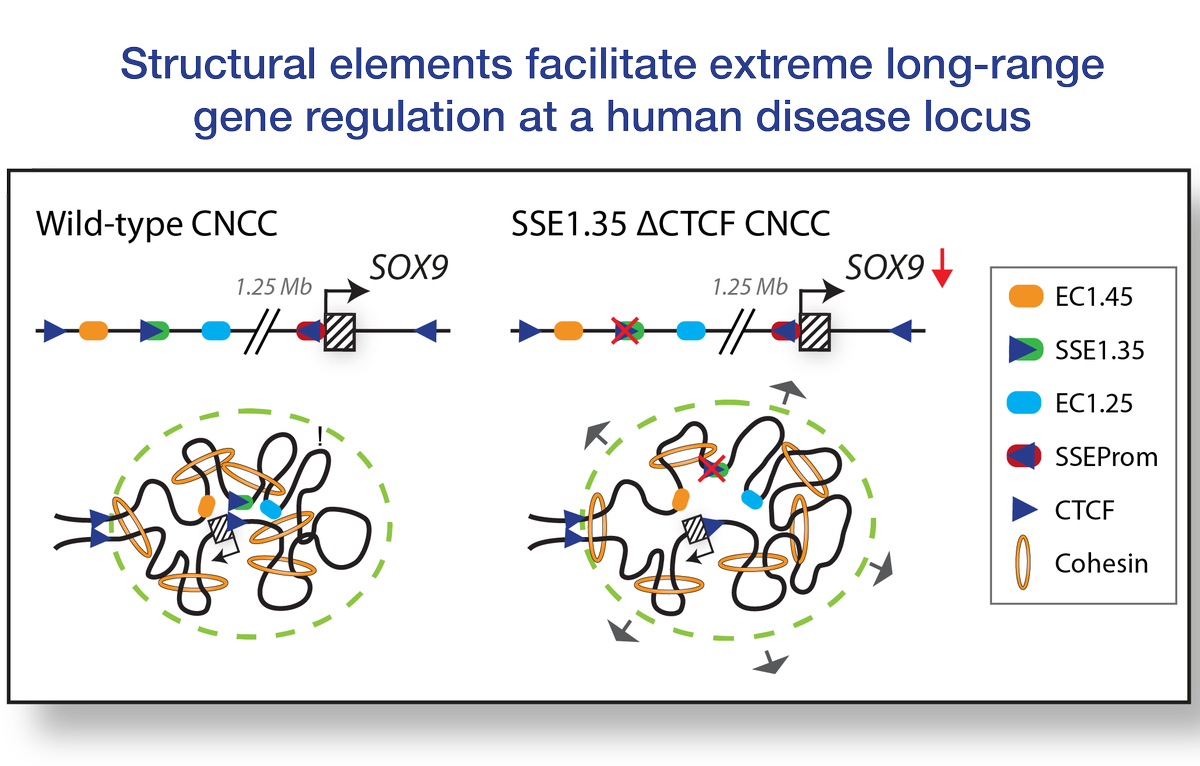
Structural elements facilitate extreme long-range gene regulation at a human disease locus biorxiv.org/cgi/content/sh… #bioRxiv
