Martin Schmitz retweetledi

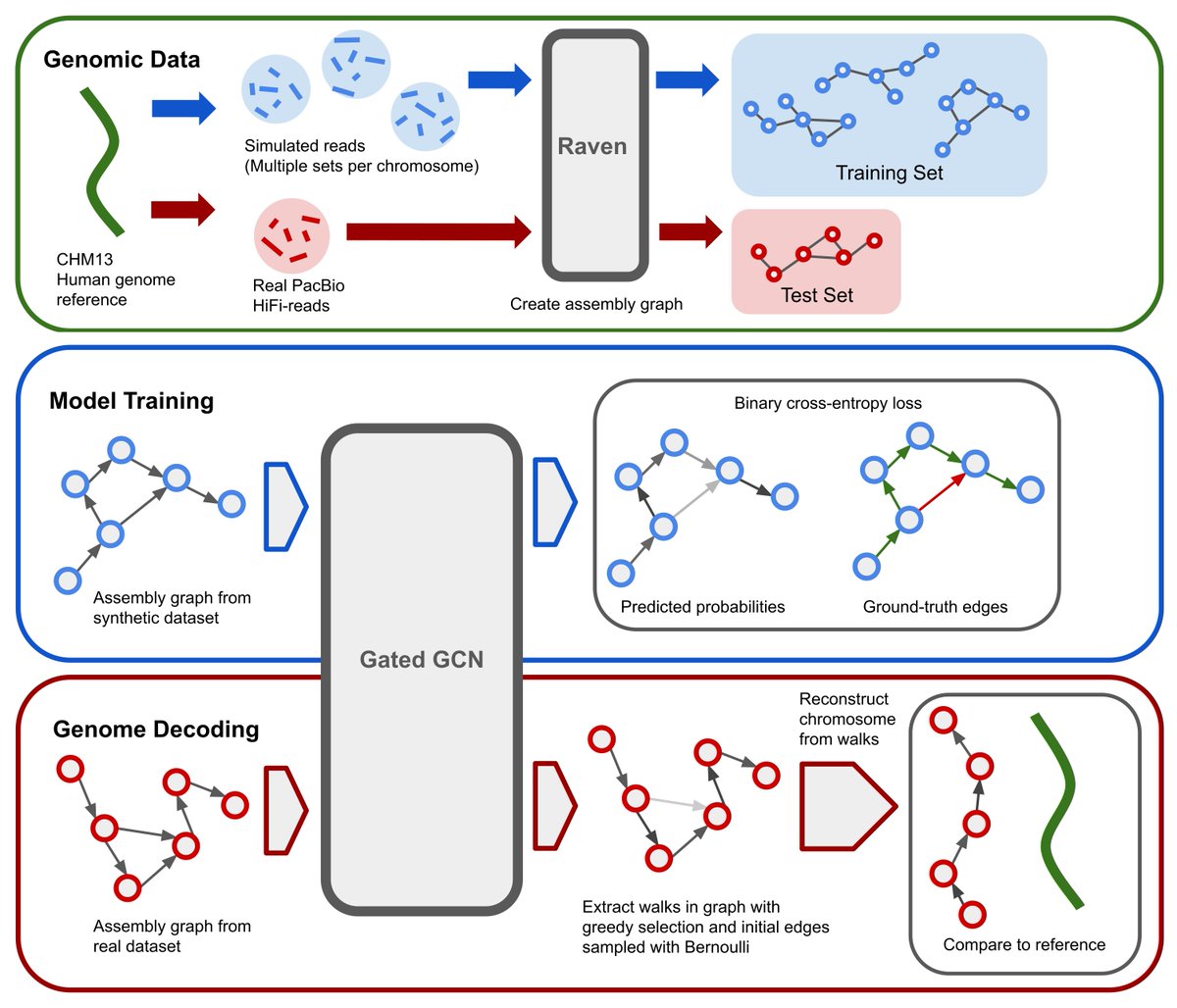
GNNome was published in @genomeresearch! This is a novel paradigm for de novo genome assembly based on GNNs. Without explicitly implementing any simplification strategies, it can achieve results comparable or higher than other SOTA tools. Paper, code, and overview are 👇 [1/8]

English