Jakob Nilsson
1.4K posts

Jakob Nilsson
@NilssonLabCph
Cell biologist working in Copenhagen. Opinions are my own
Katılım Eylül 2017
236 Takip Edilen1.1K Takipçiler

Happy to see our work on defining a dependency map of short linear motifs out. We use base editing screens to install more than 80000 mutations into SLiMs in more than 4000 proteins to determine their contribution to cell fitness. nature.com/articles/s4159…
English

Looking forward to a great line up of speakers at #CRISPRMED26 in Copenhagen: linkedin.com/pulse/crisprme…
Come join us for some excellent science in the heart of Copenhagen - still places available
English
Jakob Nilsson retweetledi
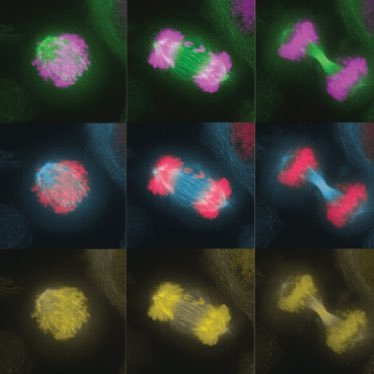
We are very excited to open the registration for the upcoming @EMBOevents on Chromosome Segregation and Aneuploidy in beautiful Stresa!!! You can find all the relevant info at meetings.embo.org/event/25-aneup…
Hope to see many of you in a few months!! 🧬🔬😍
#ChromoPloidy25
English

@LindorffLarsen A positively charged name! The alternative is more negatively charged.
English
Jakob Nilsson retweetledi

The deadline for applications is approaching!!!
Guillermo Montoya@g1_guillermo
Dear colleagues, we have a postdoctoral opening financed by the ERC_AdG project INTETOOLS @ERC_Research. Retweets much appreciated !!! see the link below for details 👇 candidate.hr-manager.net/ApplicationIni… Deadline for applications 6-11-24
English
Jakob Nilsson retweetledi
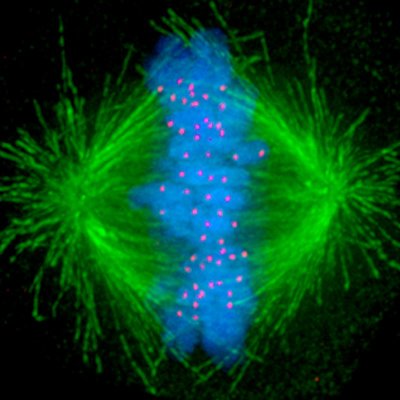
Our latest work from lab member jimmy Ly! This fundamentally changed the way we see mitosis. Massive rewiring of translational control. There is something for everyone and an amazing mechanism. Check it out.
Whitehead Institute@WhiteheadInst
Cells make variants of thousands of proteins. These variants are not produced indiscriminately, but rather through precise regulatory mechanisms that can meet rapidly changing needs of the cell. New work from @iaincheeseman's lab in @Nature: wi.mit.edu/news/elegant-s… @MITBiology
English
Jakob Nilsson retweetledi

We (@alexholehouse and @ProteinMagnus) started IDPSeminars in May 2020. It has been a wonderful platform to host an incredible array of international science. What began as a pandemic alternative to conferences has blossomed into a consistent and supportive community.
English

@lapassmore @MRC_LMB @ERC_Research @RappsilberLab @FJ_OReilly @giselleclee nice - we also found a CCR4-Not motif in ANKRD17 in our latest work with @DaveyLab. Look forward to reading this work more carefully!
English

New work from the lab!
James Stowell led this project showing the importance of multivalency and phospho-regulation in mRNA decay. This has many parallels with other mechanisms in gene expression... 🧵 1/
#RNA #NMR #bioRxiv @MRC_LMB @ERC_Research

English

Still buzzing after @duxin_lab retreat!
Feeling energised after three days of full of inspiration, fresh perspectives, and some epic fun (gokarts included!).
Thanks @NilssonLabCph for joining us on the last day, we'll get the cup back!
English
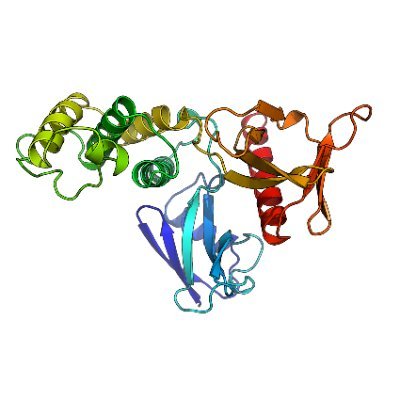
Just a few hours into the new year, and the first paper of the year involving our lab has been published🥳
Read about the substrate specificity of the B55 subunit of PP2A and its inhibitor we designed.
Great collaboration with @NilssonLabCph, @KettenbachLab and @torbenheick labs!
English

Happy to have been a part of this collaboration with @NilssonLabCph @OraFurman @KettenbachLab @jp_arulanandam. Nice to see it finally out!
Substrate recognition principles for the PP2A-B55 protein phosphatase | Science Advances science.org/doi/10.1126/sc…
English

@wgarland10 @OraFurman @KettenbachLab @jp_arulanandam Thanks for a great collaboration Will and Torben!
English

Our paper on PP2A-B55 substrate recognition out now. Great collaboration with @FurmanLab @torbenheick @KettenbachLab @jp_arulanandam. We integrate AlphaFold modelling and protein design with biochemical and cell based assays
science.org/doi/10.1126/sc…
English
