Omics Discovery retweetledi
Omics Discovery
251 posts

Omics Discovery retweetledi

Moving forward with the idea of proteomics pipelines in the cloud, collaboration with @OpenMSTeam. Label-free pipelines in development using @nextflowio @nf_core @BioContainers @OpenMSTeam #MSstats #SDRFproteomics

English
Omics Discovery retweetledi

Thanks to @JohnRYatesIII and @JProteomeRes for the invitation to publish a perspective on sample metadata in public repositories pubs.acs.org/doi/abs/10.102… , Thanks to all @EuBIC_ms contributors. Since the publication, we have opened 10 new PRs in the repo github.com/bigbio/proteom…
English
Omics Discovery retweetledi

About half of #systemsbiology models fail to directly reproduce #simulation results! Over the past 3 years we investigated >450 models published in 152 journals & identified the root cause for the lack of #reproducibility. Read our #biorxiv_sysbio article doi.org/10.1101/2020.0…
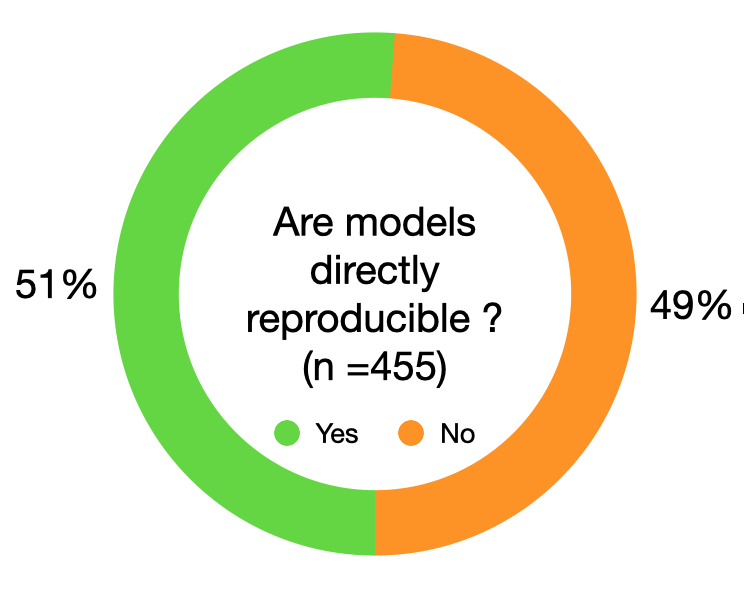
English
Omics Discovery retweetledi

Sample metadata in Proteomics repositories youtube.com/watch?v=ybueEa… github.com/bigbio/proteom… a project from @EuBIC_ms
YouTube
Català
Omics Discovery retweetledi
Omics Discovery retweetledi

The @BioContainers Registry is a light-metadata interface to quickly find the tool and the corresponding container (biocontainers.pro/#/registry). You can get info about NAME, Number of downloads, short description, License. What other information you would like to see?

English
Omics Discovery retweetledi

For those that would like to know more about SDRF here the link to the @EuBIC_ms Hackathon eventbrite.co.uk/e/eubic-hackat…
RT !!!! #ASMS2020 #proteomics @ProteomeXchange
English
Omics Discovery retweetledi

You can register for the Hackathon next week on Sample metadata annotations hosted by @EuBIC_ms.
Yasset Perez-Riverol@ypriverol
Online Hackathon and Workshop about the sample to data file format (SDRF) in proteomics. @EuBIC_ms #HUPOPSI @pride_ebi @ExpressionAtlas eventbrite.co.uk/e/eubic-hackat….
English
Omics Discovery retweetledi

If you are wondering how to sample annotations will look like in Proteomics. Here an example of the first annotated submission in @pride_ebi with Sample annotations from @GygiLab ebi.ac.uk/pride/archive/… . All the annotations have been done based on SDRF (github.com/bigbio/proteom…)
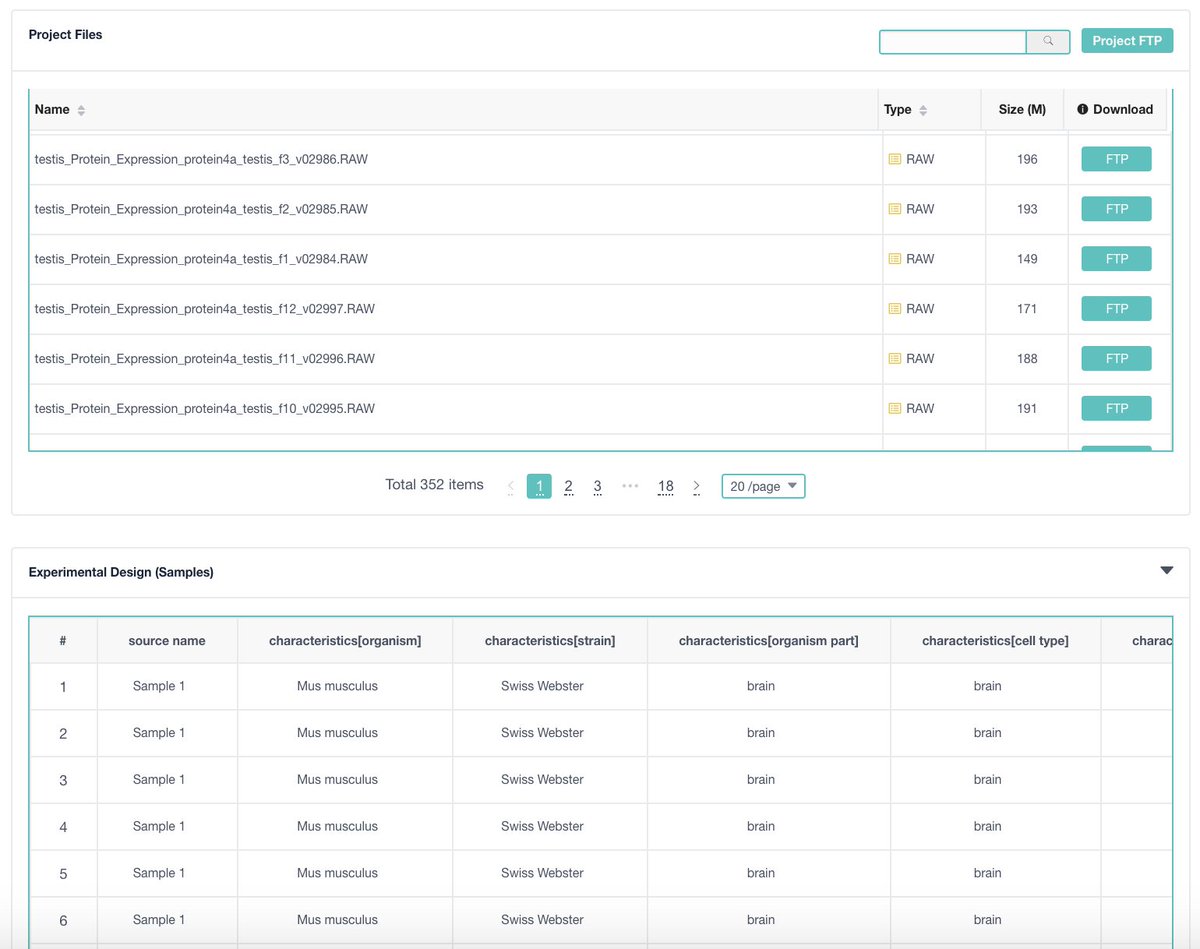
English
Omics Discovery retweetledi

Online Hackathon and Workshop about the sample to data file format (SDRF) in proteomics. @EuBIC_ms #HUPOPSI @pride_ebi @ExpressionAtlas eventbrite.co.uk/e/eubic-hackat….

English
Omics Discovery retweetledi
Omics Discovery retweetledi

Thanks to @EuBIC_ms the new SDRF file format for proteomics is really getting adopted. Multiple contributions from @EuBIC_ms members such as @mvaudel @OpenMSTeam @lgatt0 and Lev Levitsky, Johannes Griss github.com/bigbio/proteom… . We hope more people get involved.
English
Omics Discovery retweetledi

Should we make the experimental design based on the SDRF (sample to a data file format - github.com/bigbio/proteom…) on each PRIDE submission:
English
Omics Discovery retweetledi

Beta version of an Annotator of the SDRF is now deployed annotator.multiomics.info semantic auto-complete of ontology terms.
English
Omics Discovery retweetledi

The specification is here (github.com/bigbio/proteom…). If you have submitted a dataset to @pride_ebi help us to re-annotate them.
English
Omics Discovery retweetledi

61 projects annotated using the SDRF specification (github.com/bigbio/proteom…). TMT, SILAC, Label-Free, phospho-proteomics, protein-protein interactions experiments. A validator is now in place, every time a collaborator add a new experiment, the PR will be validate before merge.
English
Omics Discovery retweetledi

Finally out: OmicsDI REST interface academic.oup.com/nar/advance-ar…
English
Omics Discovery retweetledi
Omics Discovery retweetledi

Great effort from @OpenMSTeam, @MuenchLab, and myself to annotated the SARS-Cov-2 submission Experimental design in SDRF github.com/bigbio/proteom…
English