Pak DNA
49 posts






@PakGenome Can you do for vellala and nagarathars?




We received Y-111 results for a Bangash with T1a. In the next months we will receive the final Big Y-700 results. But we can already predict that he is under yfull.com/tree/T-Y19169/ This is a T1a clade associated with Central Asia ultimately




@PakGenome @wagarrwal @AnonKing02 @1kjhgfdsa1 @Wusungk @PashtunSecular Out of the PJL.DG samples, HG02597 is Gujjar. He scores autosomally like one & on yfull shares his terminal subclade with two other Gujjars. yfull.com/tree/R-C131139/ A similar case for Kurram sample, HGDP00248, autosomally different, but subclade same yfull.com/tree/L-MF68713/


@PakGenome Can you do it for punjabi brahmin private sample


One of the most Zagros shifted pop in South Asia are Gujjars and they get around 35% Zagros Zagros N: 35.5% AASI: 24.2% EHG: 11.3% WSHG: 7.39% ANF: 12.2% CHG: 9.51 Pvalue: 0.069

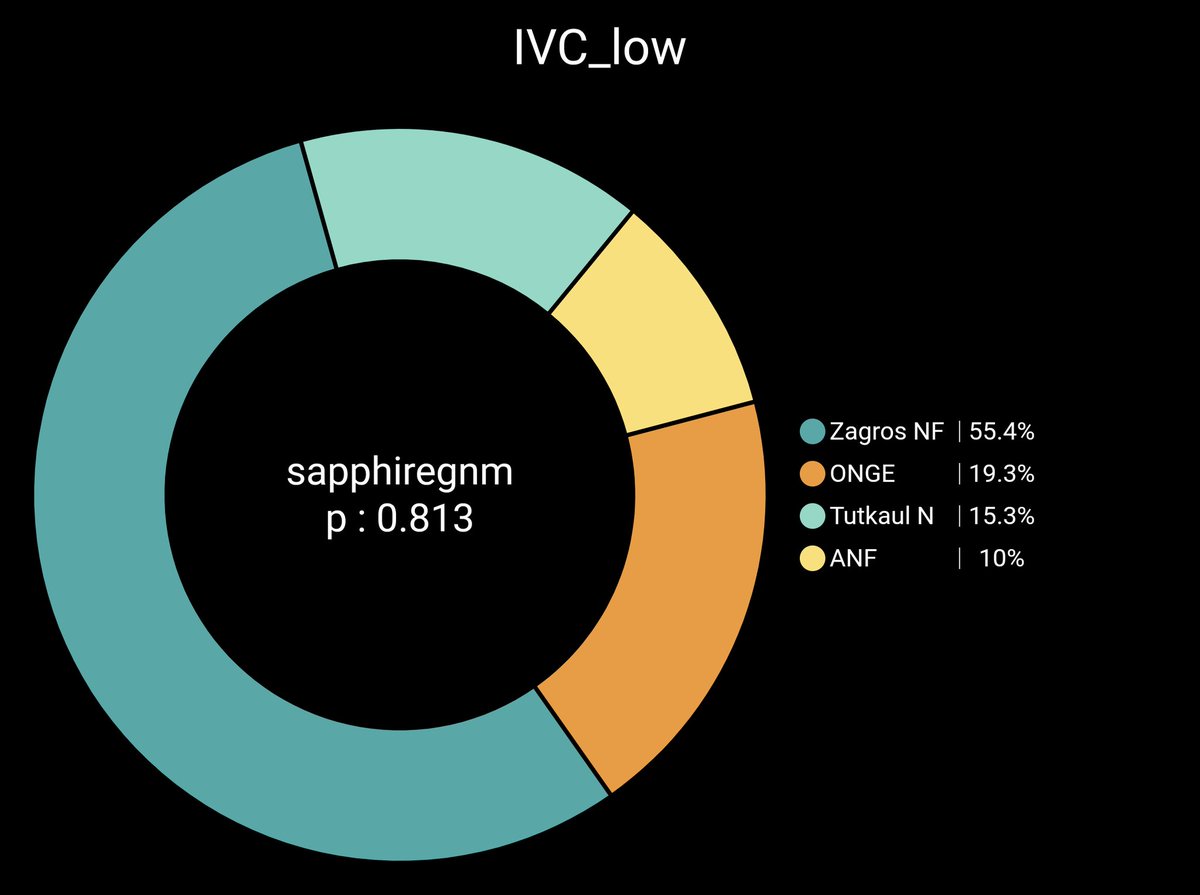


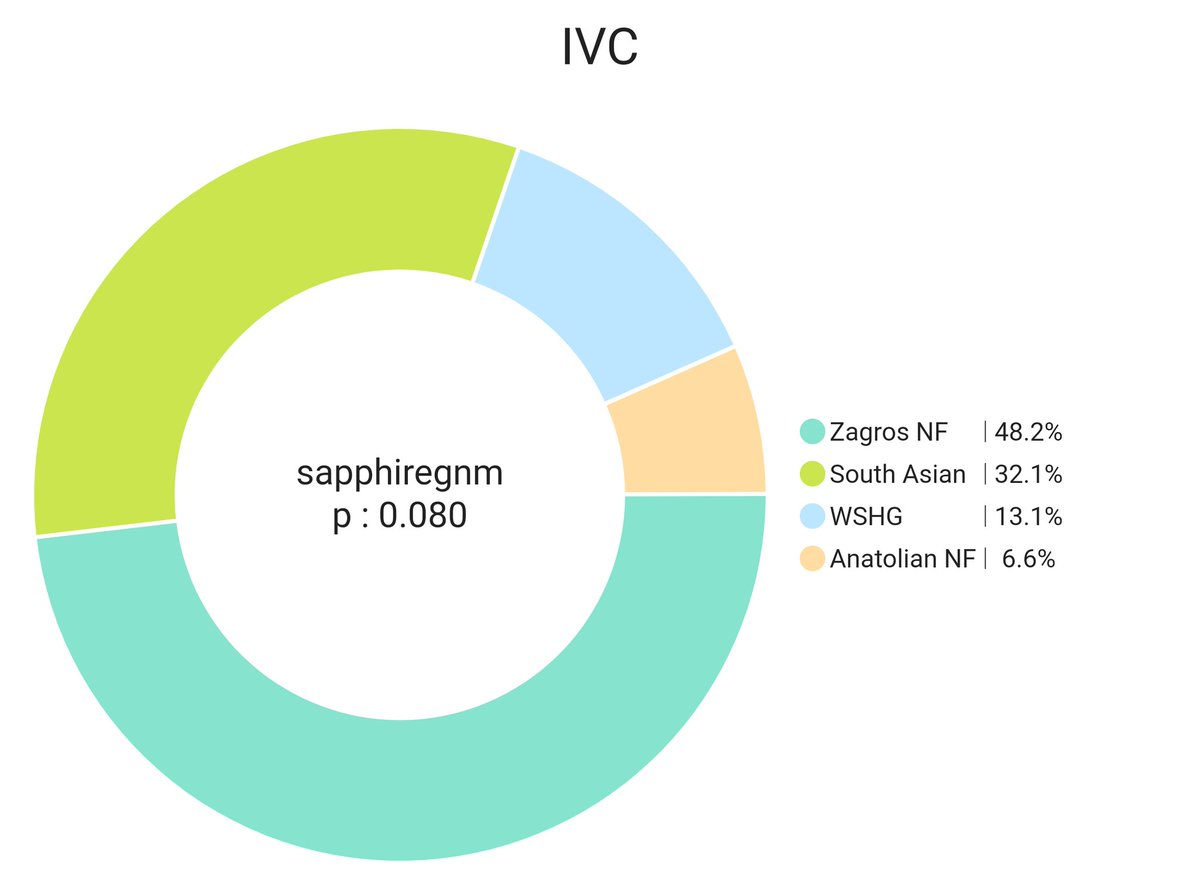

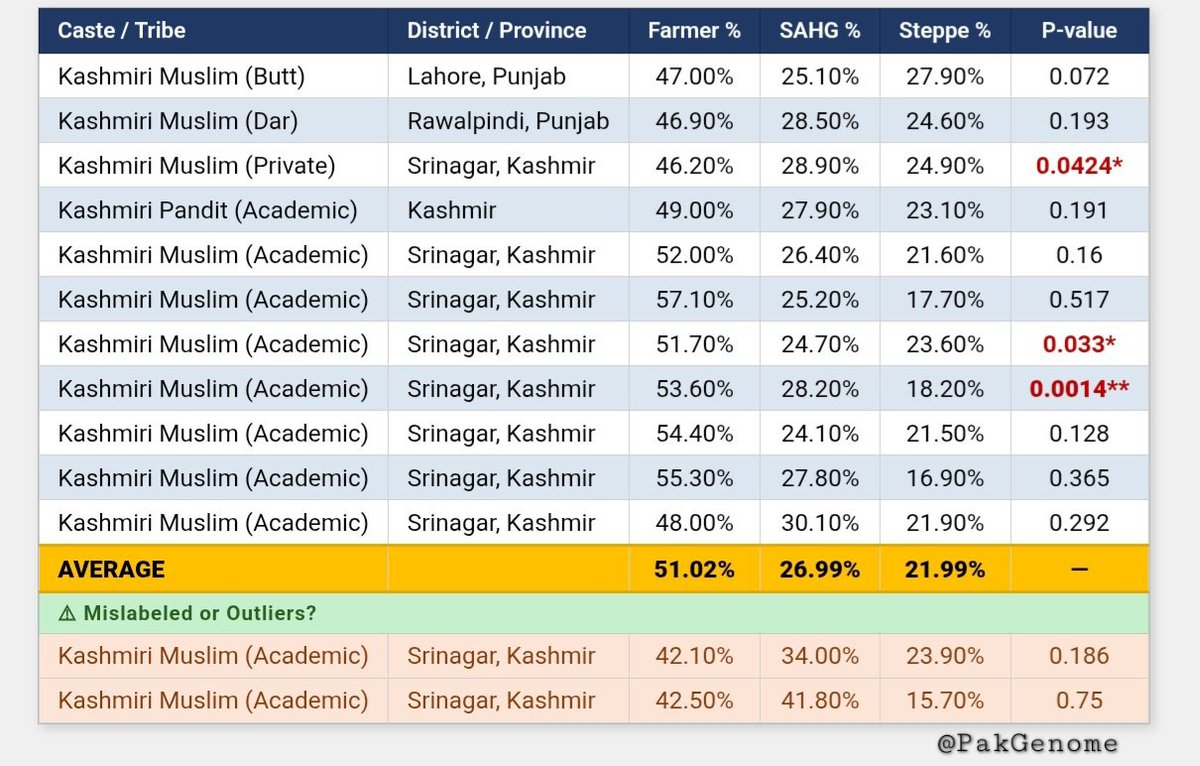
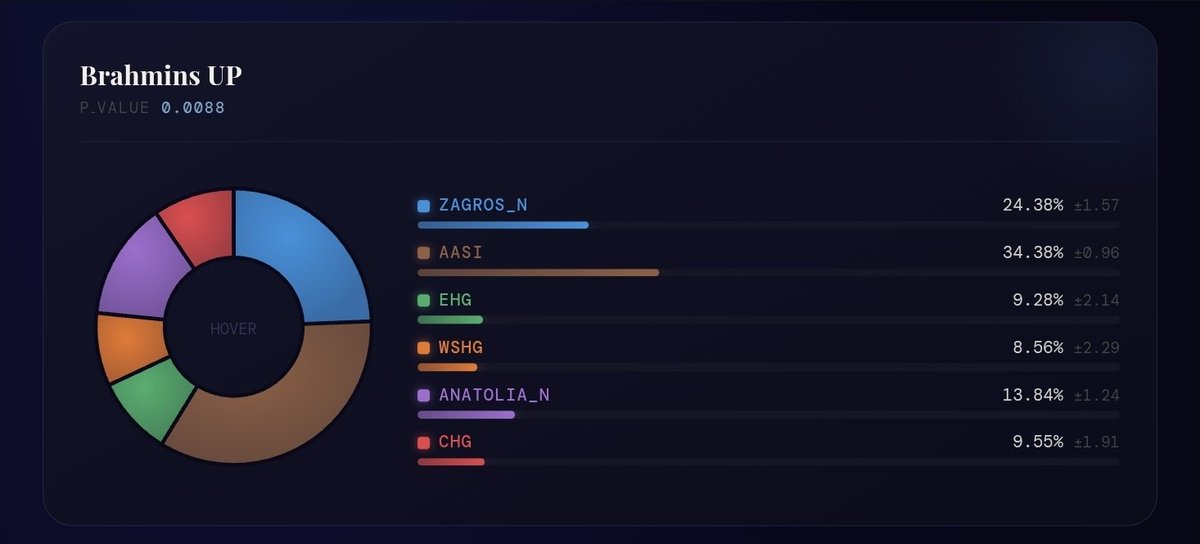
Gangetic upper castes are around 25-28% Zagros. Total farmer ancestry around 35-40%. Punjabi/Sindhi are around 30-35% Zagros. Total farmer 50-55% Total farmer is higher becoz of excess ANF, WSHG & CHG from non-Steppe sources like Indus or even Sehgabi/BMAC in case of NW groups.


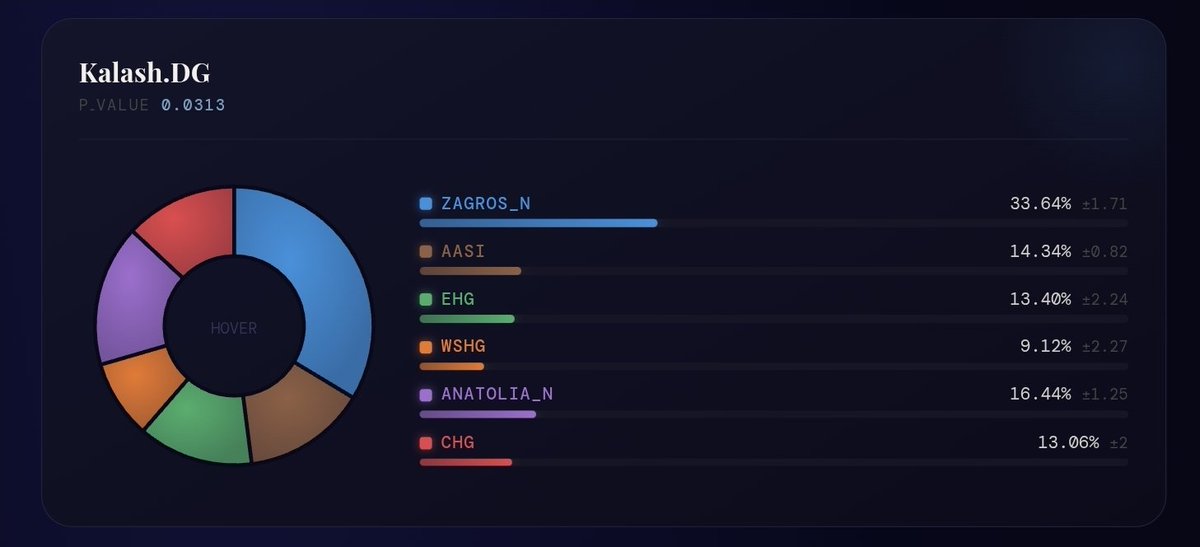
Neolithic/HG QpAdm run for Punjabi_Lahore_A samples which are a good representative of average Biradri Punjabis. Punjabis are around 30-35% Zagros. Sindhis, Pashtuns, Kohistanis and Kalash are in same range 30-35%.