

PantheonOS
9 posts

@PantheonOS
Open source AgentOS for data science @Stanford






Pantheon AI Agent Store is now live with 1300 biomedical Skills and more! Pantheon Store is a new marketplace for biomedical AI Agents, Teams, and Skills. We launch the Store with 1300+ curated bio/medical AI capabilities, built by the incredible builders behind Claude Scientific Skills, ClawBio, OpenClaw Medical Skills, LabClaw, and PantheonOS ecosystem. You can install instantly in PantheonOS (UI or CLI) and build powerful scientific workflows right now. What you can do with Pantheon Store? It can: • Discover Agents, Teams, and Skills for genomics (especially single-cell and spatial genomics), pharmacology, medicine, and bioinformatics • Install Skills seamlessly into your existing workflows • Share your own Agents, Teams, and Skills with the community Thus, Pantheon-Store turns PantheonOS into a living ecosystem for scientific AI tools. We welcome you join the community! Upload your own components. Build together. Accelerate biomedical discovery!

Pantheon AI Agent Store is now live with 1300 biomedical Skills and more! Pantheon Store is a new marketplace for biomedical AI Agents, Teams, and Skills. We launch the Store with 1300+ curated bio/medical AI capabilities, built by the incredible builders behind Claude Scientific Skills, ClawBio, OpenClaw Medical Skills, LabClaw, and PantheonOS ecosystem. You can install instantly in PantheonOS (UI or CLI) and build powerful scientific workflows right now. What you can do with Pantheon Store? It can: • Discover Agents, Teams, and Skills for genomics (especially single-cell and spatial genomics), pharmacology, medicine, and bioinformatics • Install Skills seamlessly into your existing workflows • Share your own Agents, Teams, and Skills with the community Thus, Pantheon-Store turns PantheonOS into a living ecosystem for scientific AI tools. We welcome you join the community! Upload your own components. Build together. Accelerate biomedical discovery!

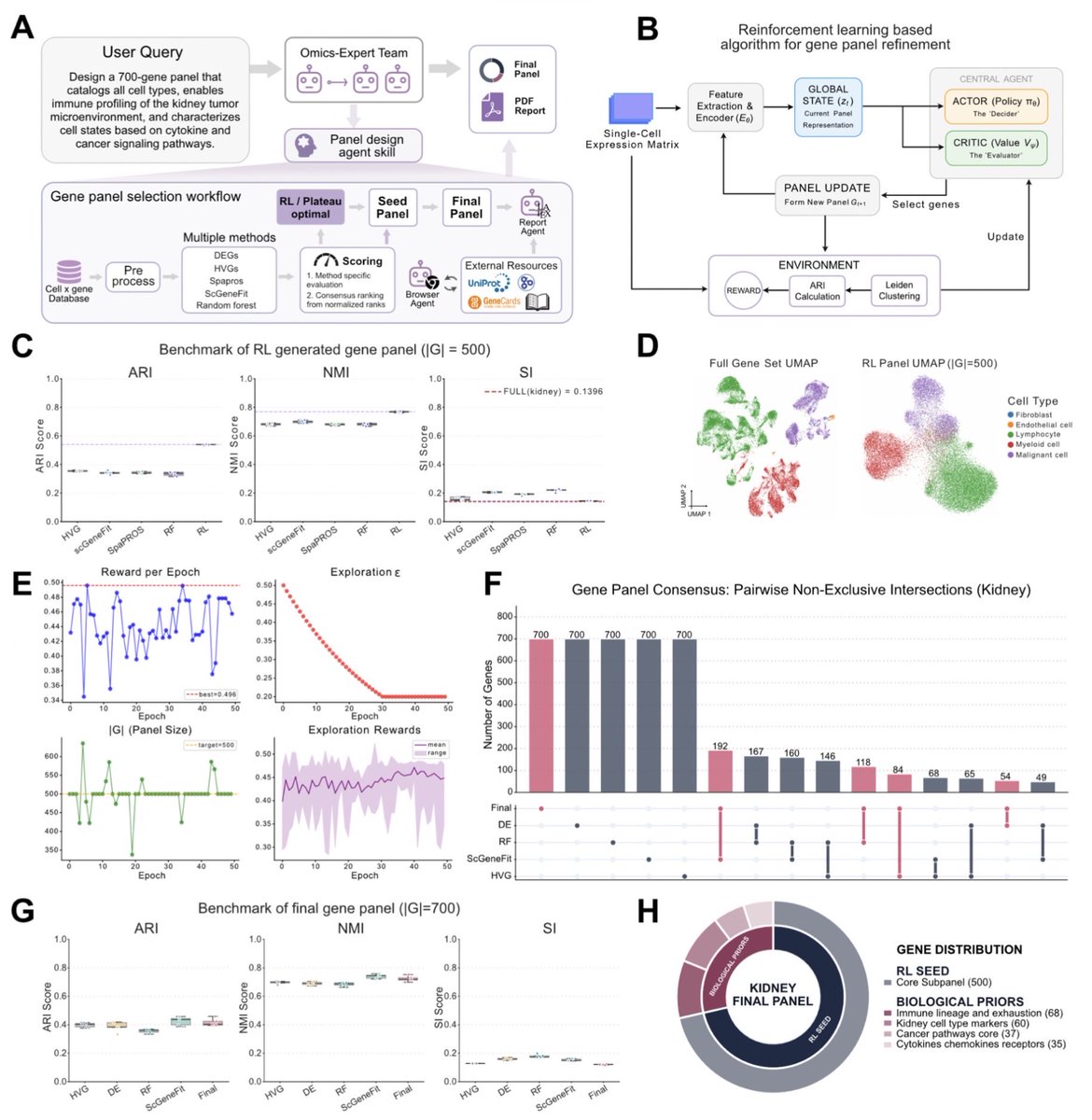
We are thrilled to share our preprint (tinyurl.com/3mtbtcwu) on PantheonOS, the first evolvable, privacy-preserving multi-agent operating system for automatic genomics discovery. 📄 Preprint: tinyurl.com/3mtbtcwu 🤖 Open-source App (free to all users): app.pantheonos.stanford.edu 🌐 More: pantheonos.stanford.edu PantheonOS unites LLM-powered agents, reinforcement learning, and agentic code evolution to push beyond routine analysis — evolving state-of-the-art algorithms to super-human performance. 🧬 Evolved batch correction (Harmony, Scanorama, BBKNN) and Reinforcement learning or RL agumented algorithms 🧠 RL–augmented gene panel design 🧭 Intelligent routing across 22+ virtual cell foundation models 🧫 Autonomous discovery from newly generated 3D early mouse embryo data 🫀 Integrated human fetal heart multi-omics with 3D whole-heart spatial data From uncovering asymmetric Cer1–Nodal inhibition in early mouse embryos to mapping spatial disease programs in the human heart, PantheonOS demonstrates a future where AI agents don’t just analyze biology — they drive discovery. This is a step toward self-evolving AI systems that accelerate science itself and push human civilization toward the singularity. This work is built over 2 years by an incredible team (Weize @Nanguage who led this project, and also to Erwin, Zhongquan @BAKEZQ, Chris @chriswzou , Zehua @starlitnightly, Yifan @YifanLu2024 , Xuehai, Zhongquan, and Miao, Cinlong's wetlab support and the entire Qiu lab. We are grateful for all our funders and industrial collaborators: @Lenovo @vizgen_inc We are excited to scale this work further and welcome philanthropic, industrial, and venture support. We invite the community (pantheonos.stanford.edu/ecosystem) to contribute, extend, and collectively reimagine the future of automated biological discovery. 🌐 More details below: