Pathogenwatch
61 posts

Pathogenwatch
@Pathogenwatch
A global platform for genomic pathogen surveillance. Delivered by @thecgps @bdi_oxford
Katılım Mart 2018
534 Takip Edilen996 Takipçiler
Pathogenwatch retweetledi

A global resource for genomic predictions of antimicrobial resistance and surveillance of Salmonella Typhi at pathogenwatch
@TheCGPS @Pathogenwatch @silargi @drkatholt @GordonDougan1 @daanensen
go.nature.com/3ivCLxQ

English
Pathogenwatch retweetledi

Se facilita la #VigilanciaGenómicaINS que sigue a la vigilancia epidemiológica. Colombia tiene información y datos abiertos para la consulta de los linajes identificados y su distribución geográfica por departamentos, disponible en micrositio #OpenDataDay #UnAñoDePandemia

Español

great to see work from @GHRUamr and WGS impact within foodborne surveillance in Latin America @NIHRglobal
@GHRUamrCol@ghruamrcol
Second PulseNetLAC day!! Brutal productive and very impresive how WGS is comming the first option in surveillance of foodborne pathogens and one health approaches in region!! Argentina, Chile, Colombia, Mexico and Perú @SomosAGROSAVIA @pahowho @MyMicroreact @Pathogenwatch
English
Pathogenwatch retweetledi

Second PulseNetLAC day!! Brutal productive and very impresive how WGS is comming the first option in surveillance of foodborne pathogens and one health approaches in region!! Argentina, Chile, Colombia, Mexico and Perú @SomosAGROSAVIA @pahowho @MyMicroreact @Pathogenwatch

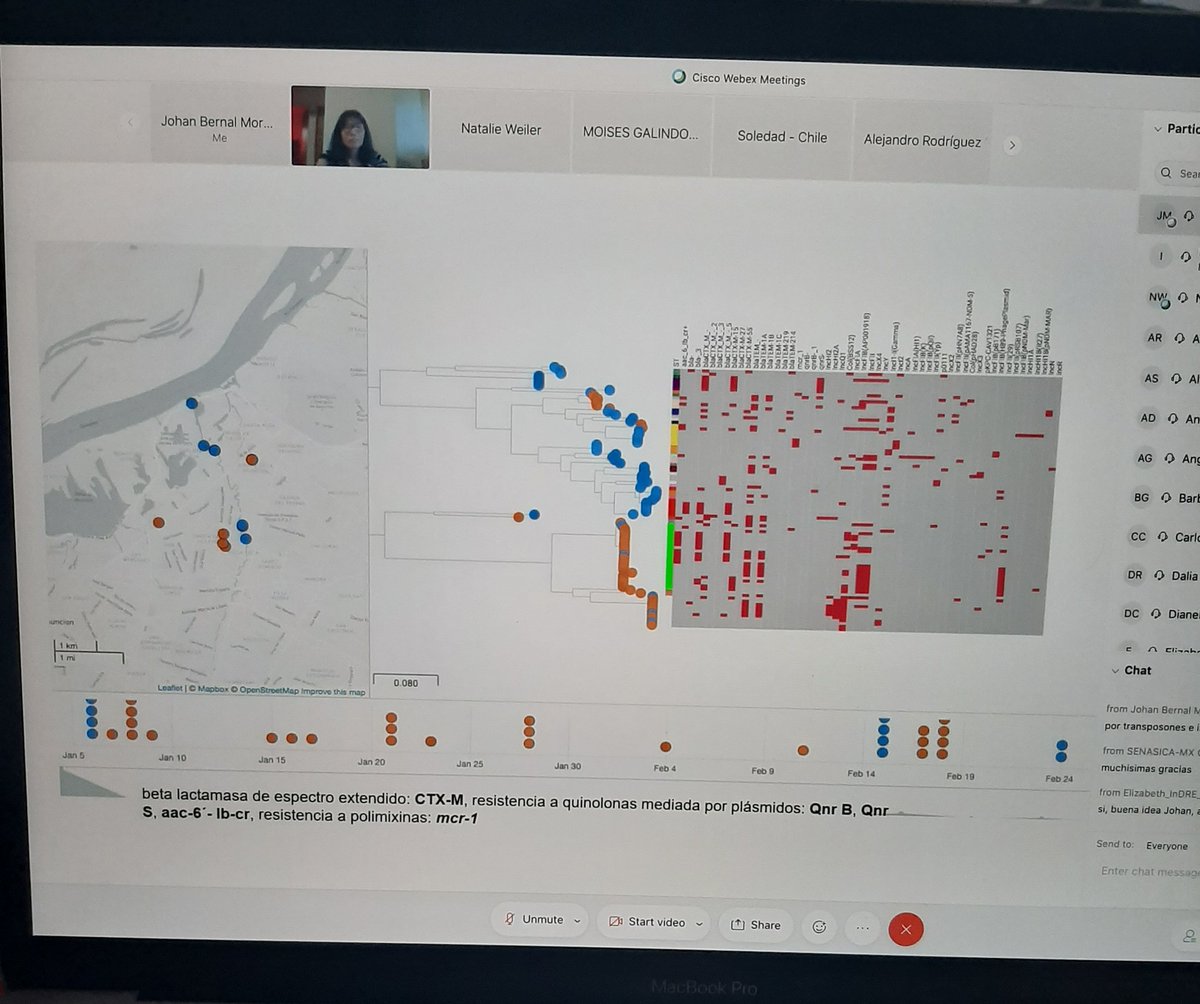
English

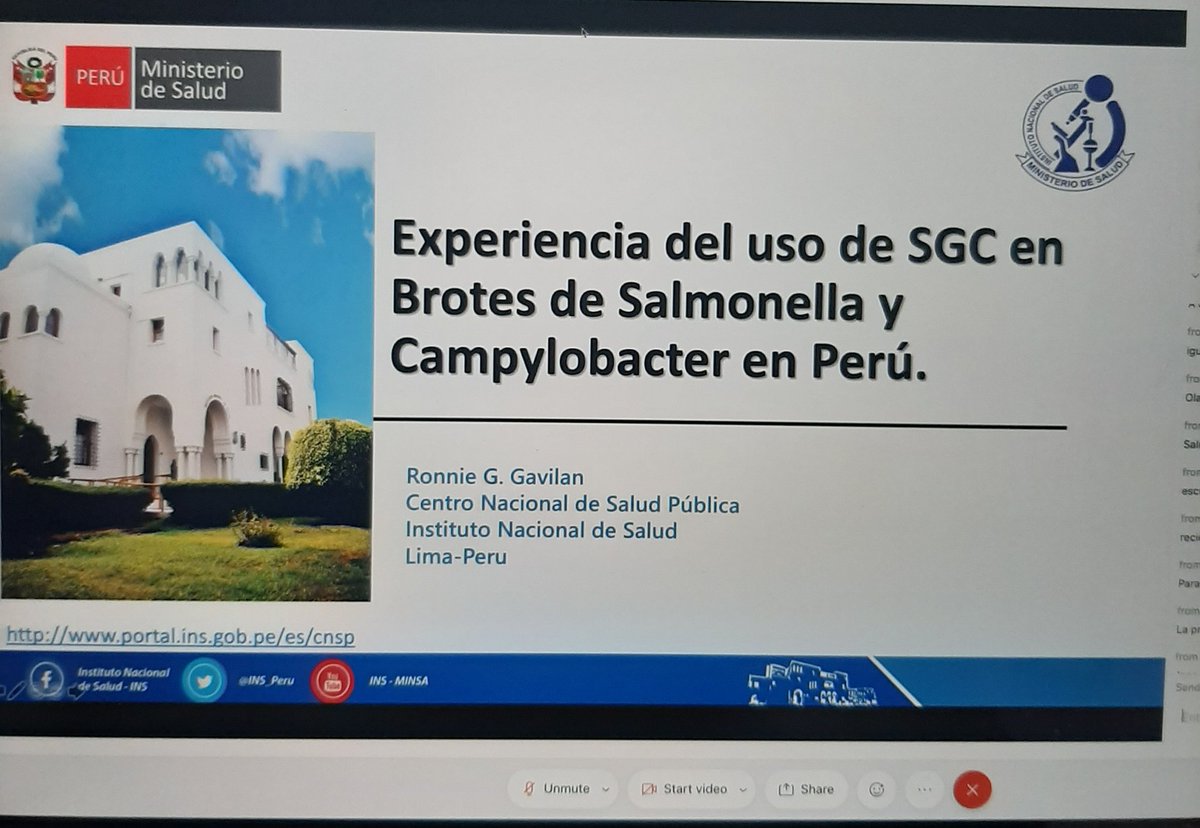
UPDATE: Addition of 937 Salmonella Typhi genomes and 7 new collections (thanks @silargi). Screenshot shows tree of 4389 public genomes with MDR strains highlighted (pathogen.watch/collection/07l…). Add your own genomes for automatic contextualisation and prediction of AMR determinants.

English

The AMR library is available at gitlab.com/cgps/pathogenw… (@leosanbu & @corinyeats). The reads were downloaded from the ENA and assembled using our pipeline: gitlab.com/cgps/ghru/pipe… (@bioinformAnt & @benmadeit). Manuscript in preparation.
English

The new AMR panel has 88 genotypes with 36 genes & >50 variants extracted from ~4000 genomes with associated MIC values and verified using Town et al (2019), Kwong et al (2018) and Yahara et al (2019). Thanks to all our collaborators including @yhgrad @thisiskatytown @gfspiteri.
English

We're excited to announce Pathogenwatch now has 14,444 N. gonorrhoeae genome assemblies (pathogen.watch/genomes/all?ge…) with location metadata and 25 new curated collections based on publications. All annotated with a new antimicrobial resistance library developed at the @TheCGPS.
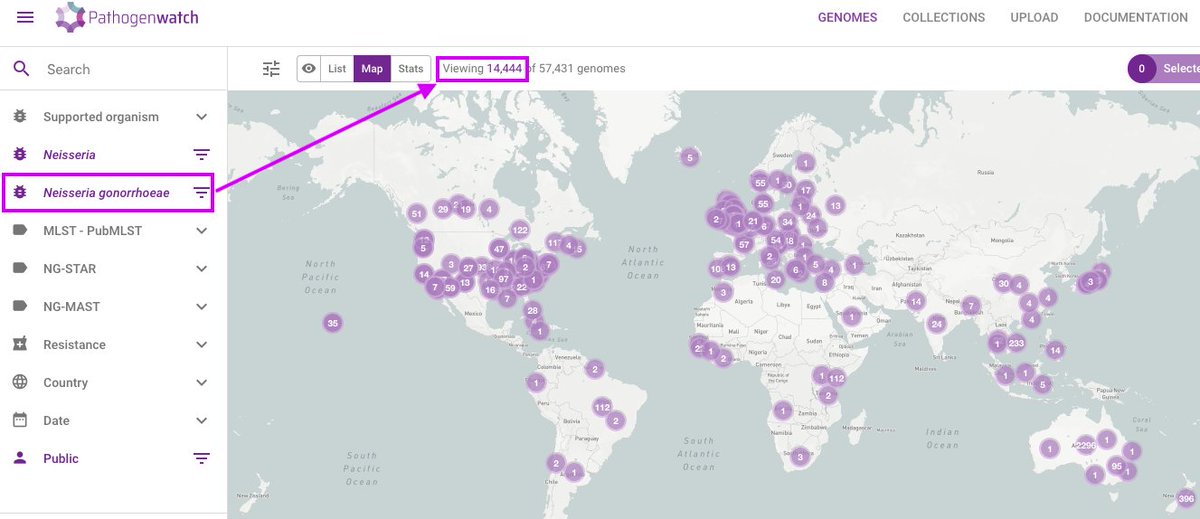
English

View the current set of genomes at pathogen.watch/genomes/all?ge….
English

We're excited to announce S. pneumoniae penicillin resistance prediction and PBP typing, along with major improvements to the current AMR library. Many thanks to Ben Metcalf and the CDC for the software and libraries and Stephanie Lo (@stephlo_lo) for validation.
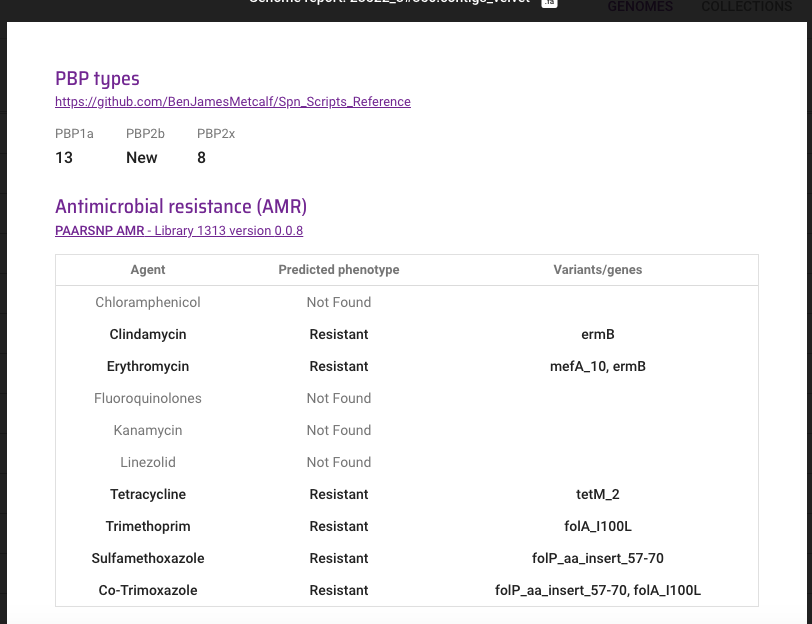
English

@DrKatHolt @pseudomonaslisa Thanks for the mention, folks. Just to clarify, we provide clustering with a network visualisation for those cgMLST schemes. We provide neighbour-joining trees for select species (see attached), with more details of the method here: cgps.gitbook.io/pathogenwatch/…
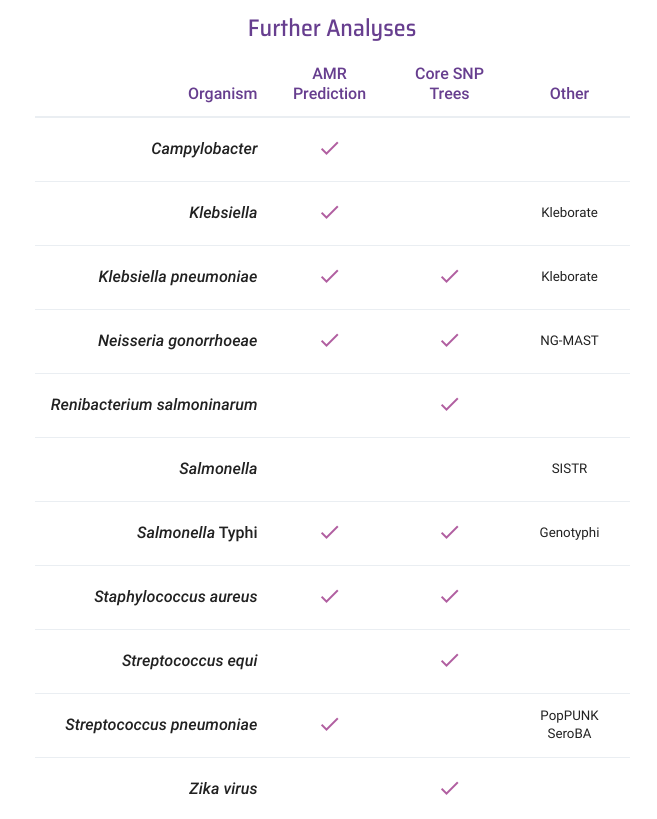
English

Trees and networks can now be exported as png or svg, and the timeline can be exported as png, for external use.
Try out the tree and timeline with datasets at pathogen.watch/collections! 🔗
(4/4)
English

Do not adjust your monitors. Pathogenwatch now has timelines! 🕰️📈
This is part of a series of upgrades designed to include functionality from @MyMicroreact, so it should feel familiar.
More details below 🧵
(1/4)

English
Pathogenwatch retweetledi