Raktim Mitra retweetledi
Raktim Mitra
85 posts


Raktim Mitra
@Raktim7879
Institute for Protein Design, University of Washington || PhD @ QCB, University of Southern California || CSE, IIT Kanpur || Soccer, Music and Painting
Los Angeles, CA Katılım Ocak 2020
142 Takip Edilen287 Takipçiler
Raktim Mitra retweetledi

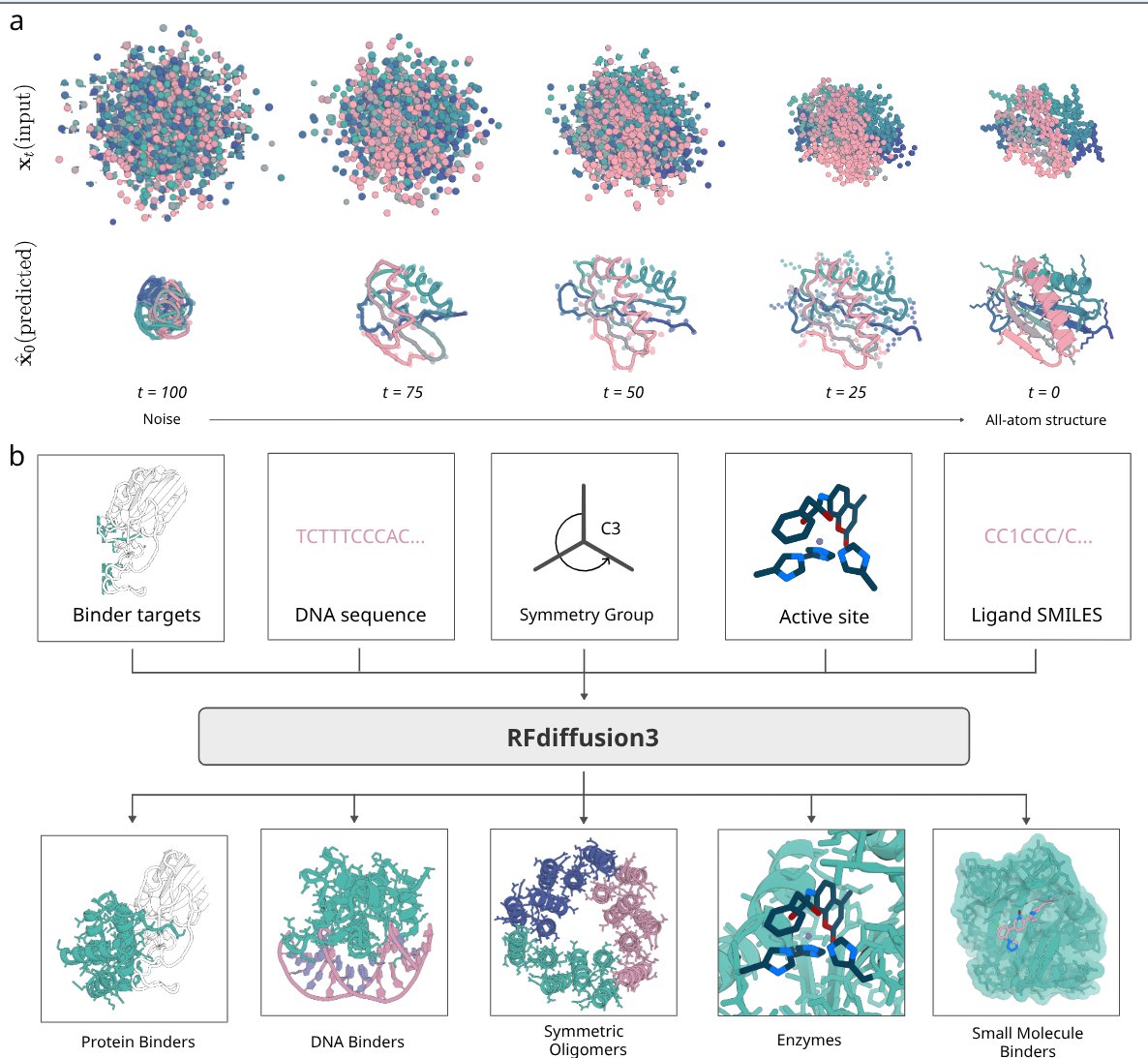
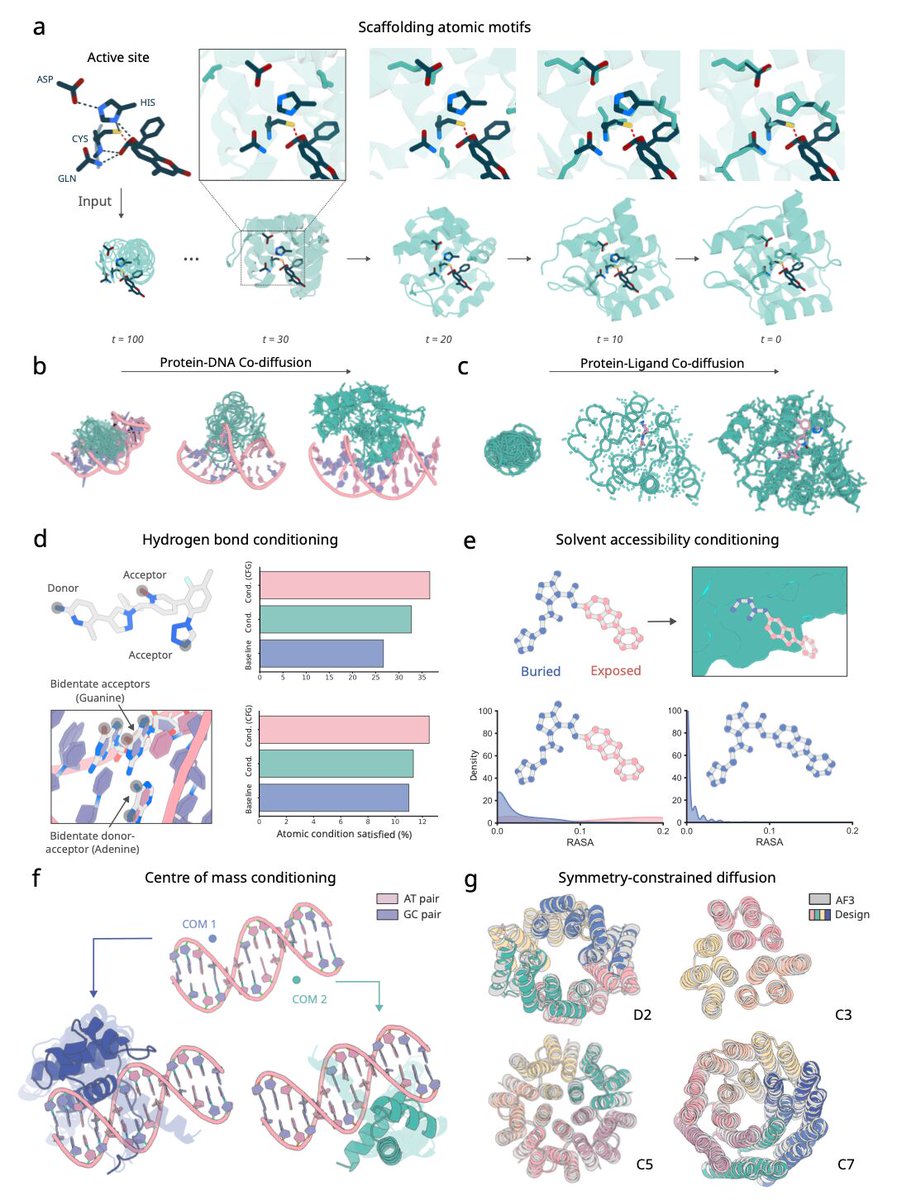
RFdiffusion3 now available! De novo protein design against any molecule
Try it on @tamarindbio today
RF3 shows success in designing de novo proteins against all-atom targets, including proteins, DNA and small molecules with diverse applications.
English

RFDiffusion3 is now open source. We took our time to make it nice and ready for you. Give it a shot 🚀 github.com/RosettaCommons…
Thanks @ichaydon for turning the DBP trajectory i generated into amazing graphic.
Thanks to @butcher_jasper @r_krishna3 and others for working with me.
Rohith Krishna@r_krishna3
Today, we report a method for design of active enzymes, RFdiffusion2, in @naturemethods. For the first time, we are able to design enzymes with native-range catalytic activity. We also are releasing our next frontier model, RFdiffusion3, code 👇
English
Raktim Mitra retweetledi

We're excited to share our new work, GUANACO! 🦙 A Python package designed for interactive single-cell data exploration and easy sharing.
GUANACO empowers users with biology-focused visualizations and flexible figure customization with clicks.
Try it here: guanaco-demo.chen-sysimeta-lab.com
bioRxiv Bioinfo@biorxiv_bioinfo
GUANACO: A Unified Web-Based Platform for Single-Cell Multi-Omics Data Visualization biorxiv.org/content/10.110… #biorxiv_bioinfo
English

RFDiffusion3 generates all atom bound conformation, making it significant for flexible targets like DNA.
An excellent teamwork to achieve something impossible by any one of us in just few months.
@butcher_jasper
@r_krishna3
biorxiv.org/content/10.110…
GIF
English
Raktim Mitra retweetledi
Raktim Mitra retweetledi

RFdiffusion3 is out! Incredible team effort led by the amazing @butcher_jasper. We've already found that RFD3 enables new classes of protein design in the lab and are excited to share our results with the world. Code coming soon!
Jasper Butcher@butcher_jasper
Very excited to share our paper "De novo Design of All-atom Biomolecular Interactions with RFdiffusion3", now on BioRXiv. biorxiv.org/content/10.110… 1/n
English
Raktim Mitra retweetledi

Very excited to share our paper "De novo Design of All-atom Biomolecular Interactions with RFdiffusion3", now on BioRXiv. biorxiv.org/content/10.110… 1/n
GIF
English
Raktim Mitra retweetledi

Accelerating Biomolecular Modeling with AtomWorks and RF3
🚀 New preprint from David Baker!🚀
1. A new framework called AtomWorks has been introduced to revolutionize biomolecular modeling. AtomWorks provides a unified and modular platform for developing state-of-the-art biomolecular models, including structure prediction, protein design, and sequence design. It streamlines the process of data preparation and model training, making it easier for researchers to prototype and test new ideas.
2. The AtomWorks framework emphasizes high-quality data handling. It standardizes inputs from diverse sources, such as the Protein Data Bank (PDB), and resolves common issues like incorrect bond orders, charges, and missing coordinates. This results in higher-quality derived features and improved model performance. For example, AtomWorks-generated reference conformers have lower energies compared to those from other open-source models.
3. AtomWorks enables rapid prototyping by breaking down data processing and featurization into modular components. This modular design allows researchers to reuse core building blocks across different networks and easily add new features. It also simplifies the integration of various datasets, facilitating the training of models like RF3 on a diverse set of biomolecular structures.
4. The framework supports scalable training of biomolecular models. AtomWorks shares most of its code across different networks, allowing researchers to repurpose existing components and improve common operations. This efficiency is demonstrated by the ability to process large batches of data quickly, such as processing a 10,000-token batch through the LigandMPNN pipeline in the time it takes for a single forward/backward pass.
5. AtomWorks is accompanied by industry-grade testing and comprehensive documentation. This ensures that the framework is reliable and easy to use, even for researchers without extensive software development experience. The documentation includes worked examples illustrating how to develop pipelines for various biomolecular modeling tasks.
6. Using AtomWorks, the authors trained RosettaFold-3 (RF3), an all-atom biomolecular structure prediction network. RF3 incorporates novel features such as implicit chirality representations and atom-level geometric conditioning, which improve its performance on tasks like predicting chiral ligands and fixed-backbone conformations.
7. RF3 simplifies dataset integration by supporting direct loading from raw crystallographic information files (CIF). The authors introduced new distillation datasets, including a nucleic acid complex distillation set and an RNA distillation set, to enhance the model's training. Additionally, RF3 includes a disordered distillation set to address issues with hallucinated secondary structures.
8. RF3 accurately adheres to specified stereochemistry out-of-the-box, without requiring inference-time guidance. It represents stereochemistry by the sign of the angles formed by atoms surrounding each chiral center and uses data augmentation techniques to improve chirality handling. As a result, RF3 predicts the correct chirality for 88% of ligand chiral centers in the test set, compared to 84% for AlphaFold3 and 76% for Boltz-2.
9. RF3 enables flexible user control through arbitrary atom-level conditioning. Users can specify distances between atoms to incorporate experimentally derived constraints, perform protein-ligand docking, or fold proteins around specific ligand conformers. This feature significantly improves the accuracy of protein-ligand interface predictions.
10. RF3 narrows the performance gap between existing open-source structure prediction models and AlphaFold3. It demonstrates competitive performance on various tasks, such as predicting protein-protein interfaces, protein-ligand interactions, and mixed L/D peptides. When trained on data up to January 2024, RF3 shows further improvements in performance.
11. The authors also trained ProteinMPNN and LigandMPNN using AtomWorks, demonstrating comparable performance to the original models. This highlights the versatility of the AtomWorks framework in supporting different biomolecular modeling tasks.
12. The AtomWorks framework and RF3 model are released with curated training data, code, and model weights, making them accessible for further research and development in the field of biomolecular modeling.
💻Code: github.com/RosettaCommons…
📜Paper: biorxiv.org/content/10.110…
#BiomolecularModeling #AtomWorks #RF3 #StructurePrediction #ProteinDesign #OpenSource #MachineLearning #DeepLearning #ComputationalBiology

English
Raktim Mitra retweetledi

AtomWorks is out! Building upon @biotite_python, we built for a toolkit for all things biomolecules and trained RF3 with it. All open-source, test it via `pip install atomworks`!
AtomWorks: github.com/RosettaCommons…
RF3: github.com/RosettaCommons…
Paper: tinyurl.com/y2w4z65b
1/6

English
Raktim Mitra retweetledi

Automated #DrugDesign with our new #GenerativeAI method. We show spatial sub-sampling enables molecular scaffold hopping, linker design, fragment growing, and pattern replacement. @JCIM_JCTC
paper: pubs.acs.org/doi/10.1021/ac…
code: github.com/jssweller/Drug…
@RemoRohs lab @qcb_usc
English
Raktim Mitra retweetledi

Excited to share our new AI drug design method DrugHIVE pubished in @JCIM_JCTC. @RemoRohs lab @qcb_usc. #compchem #machinelearning
Structure-Based Drug Design with a Deep Hierarchical Generative Model
pubs.acs.org/doi/10.1021/ac…
GitHub: github.com/jssweller/Drug…
🧵1/6
English
Raktim Mitra retweetledi

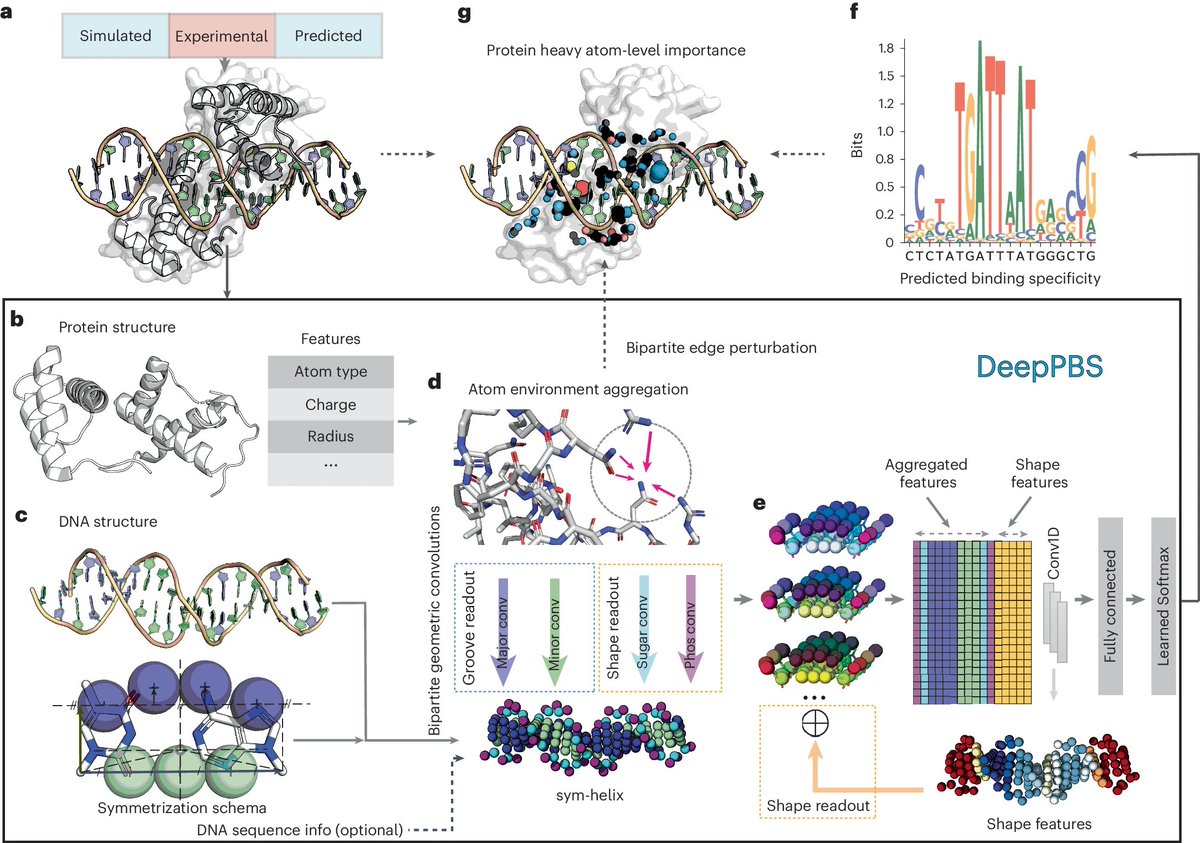
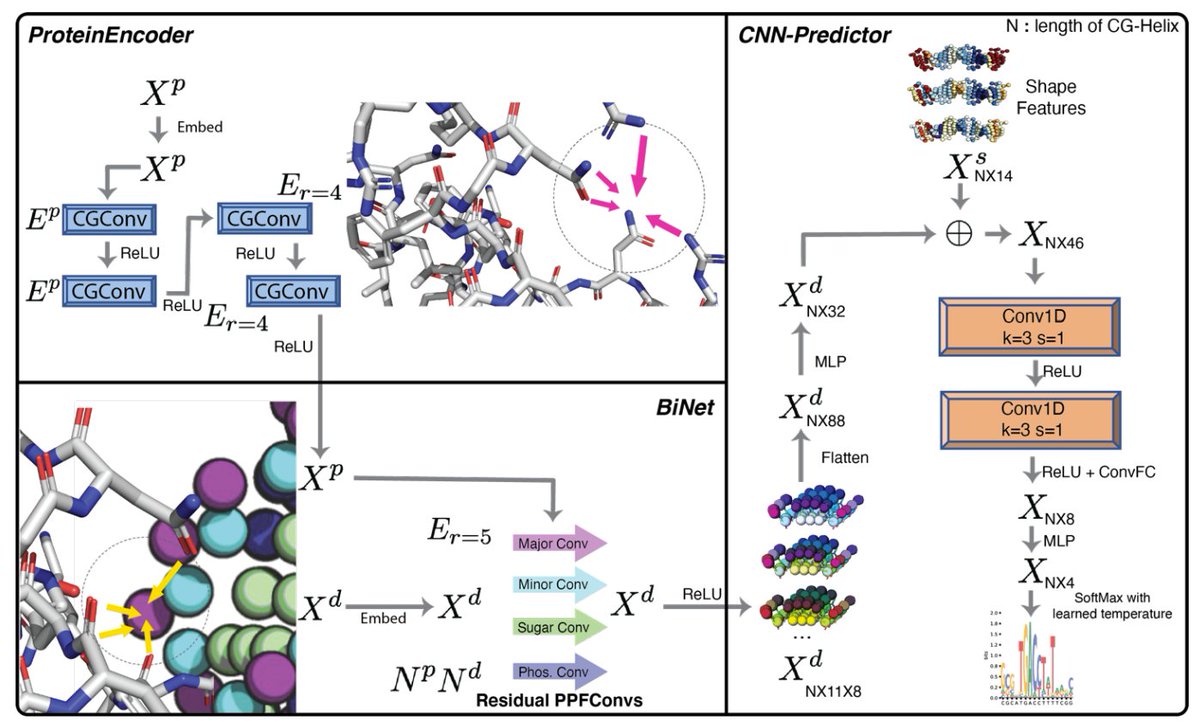
Geometric deep learning of protein–DNA binding specificity | @naturemethods
- Represent DNA structures as symmetrized helices, and Proteins as atom-based graphs with (one-hot atom type, solvent-accessible surface feature, and Atchley factors etc.)
- Use spatial Graph Convolution, Bipartite Geometric Network (BiNet), and CNN-Predictor Modules for specificity prediction
- Separate convolutions applied for major groove, minor groove, phosphate, and sugar moieties
- Train on binding specificity data from JASPAR and HOCOMOCO databases
Link: nature.com/articles/s4159…
English
Raktim Mitra retweetledi

Geometric deep learning of protein–DNA binding specificity
nature.com/articles/s4159…
Brisbane, Queensland 🇦🇺 English
Raktim Mitra retweetledi